SigD
- Description: RNA polymerase sigma factor SigD
| Gene name | sigD |
| Synonyms | flaB |
| Essential | no |
| Product | RNA polymerase sigma factor SigD |
| Function | regulation of flagella, motility, chemotaxis and autolysis |
| MW, pI | 29 kDa, 5.167 |
| Gene length, protein length | 762 bp, 254 aa |
| Immediate neighbours | cheD, swrB |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 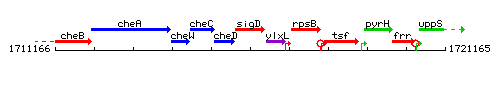
This image was kindly provided by SubtiList
| |
Contents
Categories containing this gene/protein
transcription, sigma factors and their control, motility and chemotaxis
This gene is a member of the following regulons
CodY regulon, SigD regulon, Spo0A regulon
The SigD regulon
The gene
Basic information
- Locus tag: BSU16470
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family: sigma-70 factor family (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Localization:
Database entries
- Structure:
- UniProt: P10726
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Operon:
- flgB-flgC-fliE-fliF-fliG-fliH-fliI-fliJ-ylxF-fliK-ylxG-flgE-fliL-fliM-fliY-cheY-fliZ-fliP-fliQ-fliR-flhB-flhA-flhF-ylxH-cheB-cheA-cheW-cheC-cheD-sigD-swrB PubMed
- ylxF-fliK-ylxG-flgE-fliL-fliM-fliY-cheY-fliZ-fliP-fliQ-fliR-flhB-flhA-flhF-ylxH-cheB-cheA-cheW-cheC-cheD-sigD-swrB PubMed
- sigD-swrB PubMed
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system: B. pertussis adenylate cyclase-based bacterial two hybrid system (BACTH), available in Stülke lab
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Reviews
The SigD regulon
Original publications
Additional publications: PubMed