Difference between revisions of "FliY"
(→Expression and regulation) |
|||
| Line 106: | Line 106: | ||
=Expression and regulation= | =Expression and regulation= | ||
| − | * '''Operon:''' | + | * '''Operon:''' part of the [[fla-che operon]] |
** ''[[flgB]]-[[flgC]]-[[fliE]]-[[fliF]]-[[fliG]]-[[fliH]]-[[fliI]]-[[fliJ]]-[[ylxF]]-[[fliK]]-[[flgD]]-[[flgE]]-[[fliL]]-[[fliM]]-[[fliY]]-[[cheY]]-[[fliZ]]-[[fliP]]-[[fliQ]]-[[fliR]]-[[flhB]]-[[flhA]]-[[flhF]]-[[flhG]]-[[cheB]]-[[cheA]]-[[cheW]]-[[cheC]]-[[cheD]]-[[sigD]]-[[swrB]]'' {{PubMed|9657996,8157612,15175317}} | ** ''[[flgB]]-[[flgC]]-[[fliE]]-[[fliF]]-[[fliG]]-[[fliH]]-[[fliI]]-[[fliJ]]-[[ylxF]]-[[fliK]]-[[flgD]]-[[flgE]]-[[fliL]]-[[fliM]]-[[fliY]]-[[cheY]]-[[fliZ]]-[[fliP]]-[[fliQ]]-[[fliR]]-[[flhB]]-[[flhA]]-[[flhF]]-[[flhG]]-[[cheB]]-[[cheA]]-[[cheW]]-[[cheC]]-[[cheD]]-[[sigD]]-[[swrB]]'' {{PubMed|9657996,8157612,15175317}} | ||
** ''[[ylxF]]-[[fliK]]-[[flgD]]-[[flgE]]-[[fliL]]-[[fliM]]-[[fliY]]-[[cheY]]-[[fliZ]]-[[fliP]]-[[fliQ]]-[[fliR]]-[[flhB]]-[[flhA]]-[[flhF]]-[[flhG]]-[[cheB]]-[[cheA]]-[[cheW]]-[[cheC]]-[[cheD]]-[[sigD]]-[[swrB]]'' {{PubMed|20233303}} | ** ''[[ylxF]]-[[fliK]]-[[flgD]]-[[flgE]]-[[fliL]]-[[fliM]]-[[fliY]]-[[cheY]]-[[fliZ]]-[[fliP]]-[[fliQ]]-[[fliR]]-[[flhB]]-[[flhA]]-[[flhF]]-[[flhG]]-[[cheB]]-[[cheA]]-[[cheW]]-[[cheC]]-[[cheD]]-[[sigD]]-[[swrB]]'' {{PubMed|20233303}} | ||
| Line 116: | Line 116: | ||
** ''[[ylxF]]'': [[SigD]] {{PubMed|20233303}} | ** ''[[ylxF]]'': [[SigD]] {{PubMed|20233303}} | ||
| − | * '''Regulation:''' | + | * '''Regulation:''' see [[fla-che operon]] |
| − | |||
| − | * '''Regulatory mechanism:''' | + | * '''Regulatory mechanism:''' see [[fla-che operon]] |
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
* '''Additional information:''' | * '''Additional information:''' | ||
Revision as of 15:16, 12 February 2015
- Description: flagellar motor switch protein
| Gene name | fliY |
| Synonyms | cheD |
| Essential | no |
| Product | flagellar motor switch protein |
| Function | movement and chemotaxis |
| Gene expression levels in SubtiExpress: fliY | |
| MW, pI | 40 kDa, 4.117 |
| Gene length, protein length | 1134 bp, 378 aa |
| Immediate neighbours | fliM, cheY |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 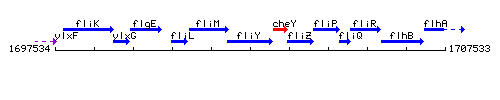
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed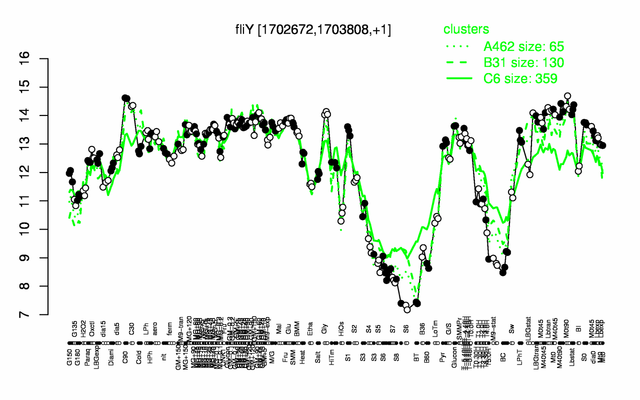
| |
Contents
Categories containing this gene/protein
motility and chemotaxis, membrane proteins, phosphoproteins
This gene is a member of the following regulons
CodY regulon, DegU regulon, SigD regulon, Spo0A regulon
The gene
Basic information
- Locus tag: BSU16320
Phenotypes of a mutant
Database entries
- BsubCyc: BSU16320
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family: fliN/mopA/spaO family (according to Swiss-Prot) cheC family
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- phosphorylated on Arg-251 PubMed
- Cofactor(s):
- Effectors of protein activity:
- Localization: cell membrane (according to Swiss-Prot)
Database entries
- BsubCyc: BSU16320
- Structure:
- UniProt: P24073
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Operon: part of the fla-che operon
- Regulation: see fla-che operon
- Regulatory mechanism: see fla-che operon
- Additional information:
- number of protein molecules per cell (minimal medium with glucose and ammonium): 1488 PubMed
- number of protein molecules per cell (complex medium with amino acids, without glucose): 1885 PubMed
- number of protein molecules per cell (minimal medium with glucose and ammonium, exponential phase): 850 PubMed
- number of protein molecules per cell (minimal medium with glucose and ammonium, early stationary phase after glucose exhaustion): 900 PubMed
- number of protein molecules per cell (minimal medium with glucose and ammonium, late stationary phase after glucose exhaustion): 713 PubMed
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Serena Mordini, Cecilia Osera, Simone Marini, Francesco Scavone, Riccardo Bellazzi, Alessandro Galizzi, Cinzia Calvio
The role of SwrA, DegU and P(D3) in fla/che expression in B. subtilis.
PLoS One: 2013, 8(12);e85065
[PubMed:24386445]
[WorldCat.org]
[DOI]
(I e)
Alexander K W Elsholz, Kürsad Turgay, Stephan Michalik, Bernd Hessling, Katrin Gronau, Dan Oertel, Ulrike Mäder, Jörg Bernhardt, Dörte Becher, Michael Hecker, Ulf Gerth
Global impact of protein arginine phosphorylation on the physiology of Bacillus subtilis.
Proc Natl Acad Sci U S A: 2012, 109(19);7451-6
[PubMed:22517742]
[WorldCat.org]
[DOI]
(I p)
Kazuo Kobayashi
Gradual activation of the response regulator DegU controls serial expression of genes for flagellum formation and biofilm formation in Bacillus subtilis.
Mol Microbiol: 2007, 66(2);395-409
[PubMed:17850253]
[WorldCat.org]
[DOI]
(P p)
H Werhane, P Lopez, M Mendel, M Zimmer, G W Ordal, L M Márquez-Magaña
The last gene of the fla/che operon in Bacillus subtilis, ylxL, is required for maximal sigmaD function.
J Bacteriol: 2004, 186(12);4025-9
[PubMed:15175317]
[WorldCat.org]
[DOI]
(P p)
Hendrik Szurmant, Travis J Muff, George W Ordal
Bacillus subtilis CheC and FliY are members of a novel class of CheY-P-hydrolyzing proteins in the chemotactic signal transduction cascade.
J Biol Chem: 2004, 279(21);21787-92
[PubMed:14749334]
[WorldCat.org]
[DOI]
(P p)
Virginie Molle, Masaya Fujita, Shane T Jensen, Patrick Eichenberger, José E González-Pastor, Jun S Liu, Richard Losick
The Spo0A regulon of Bacillus subtilis.
Mol Microbiol: 2003, 50(5);1683-701
[PubMed:14651647]
[WorldCat.org]
[DOI]
(P p)
Hendrik Szurmant, Michael W Bunn, Vincent J Cannistraro, George W Ordal
Bacillus subtilis hydrolyzes CheY-P at the location of its action, the flagellar switch.
J Biol Chem: 2003, 278(49);48611-6
[PubMed:12920116]
[WorldCat.org]
[DOI]
(P p)
W Estacio, S S Anna-Arriola, M Adedipe, L M Márquez-Magaña
Dual promoters are responsible for transcription initiation of the fla/che operon in Bacillus subtilis.
J Bacteriol: 1998, 180(14);3548-55
[PubMed:9657996]
[WorldCat.org]
[DOI]
(P p)
L M Márquez-Magaña, M J Chamberlin
Characterization of the sigD transcription unit of Bacillus subtilis.
J Bacteriol: 1994, 176(8);2427-34
[PubMed:8157612]
[WorldCat.org]
[DOI]
(P p)
D S Bischoff, G W Ordal
Identification and characterization of FliY, a novel component of the Bacillus subtilis flagellar switch complex.
Mol Microbiol: 1992, 6(18);2715-23
[PubMed:1447979]
[WorldCat.org]
[DOI]
(P p)