Difference between revisions of "RibD"
| Line 80: | Line 80: | ||
* '''Swiss prot entry:''' [http://www.uniprot.org/uniprot/P17618 P17618] | * '''Swiss prot entry:''' [http://www.uniprot.org/uniprot/P17618 P17618] | ||
| − | * '''KEGG entry:''' [http://www.genome.jp/dbget-bin/www_bget?bsu | + | * '''KEGG entry:''' [http://www.genome.jp/dbget-bin/www_bget?bsu:BSU23280] |
* '''E.C. number:''' [http://www.expasy.org/enzyme/3.5.4.26 3.5.4.26] | * '''E.C. number:''' [http://www.expasy.org/enzyme/3.5.4.26 3.5.4.26] | ||
Revision as of 05:10, 25 June 2009
- Description: 5-amino-6-(5-phosphoribosylamino)uracil reductase
| Gene name | ribD |
| Synonyms | ribG |
| Essential | no |
| Product | 5-amino-6-(5-phosphoribosylamino)uracil reductase |
| Function | riboflavin biosynthesis |
| MW, pI | 39 kDa, 6.079 |
| Gene length, protein length | 1083 bp, 361 aa |
| Immediate neighbours | ribE, ypuE |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 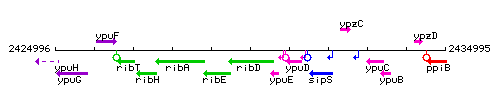
This image was kindly provided by SubtiList
| |
Contents
The gene
Basic information
- Locus tag: BSU23280
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: 2,5-diamino-6-hydroxy-4-(5-phosphoribosylamino)pyrimidine + H2O = 5-amino-6-(5-phosphoribosylamino)uracil + NH3 (according to Swiss-Prot)
- Protein family:
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Interactions:
- Localization:
Database entries
- Structure: 3EX8
- Swiss prot entry: P17618
- KEGG entry: [3]
- E.C. number: 3.5.4.26
Additional information
Expression and regulation
- Regulatory mechanism: FMN-box: riboswitch, mediates termination/ antitermination control of the operon, in the absence of FMN: antitermination, in the presence of FMN: termination PubMed
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
J Kenneth Wickiser, Wade C Winkler, Ronald R Breaker, Donald M Crothers
The speed of RNA transcription and metabolite binding kinetics operate an FMN riboswitch.
Mol Cell: 2005, 18(1);49-60
[PubMed:15808508]
[WorldCat.org]
[DOI]
(P p)
Wade C Winkler, Smadar Cohen-Chalamish, Ronald R Breaker
An mRNA structure that controls gene expression by binding FMN.
Proc Natl Acad Sci U S A: 2002, 99(25);15908-13
[PubMed:12456892]
[WorldCat.org]
[DOI]
(P p)
V N Mironov, A S Kraev, M L Chikindas, B K Chernov, A I Stepanov, K G Skryabin
Functional organization of the riboflavin biosynthesis operon from Bacillus subtilis SHgw.
Mol Gen Genet: 1994, 242(2);201-8
[PubMed:8159171]
[WorldCat.org]
[DOI]
(P p)
V Azevedo, A Sorokin, S D Ehrlich, P Serror
The transcriptional organization of the Bacillus subtilis 168 chromosome region between the spoVAF and serA genetic loci.
Mol Microbiol: 1993, 10(2);397-405
[PubMed:7934830]
[WorldCat.org]
[DOI]
(P p)