Difference between revisions of "RocA"
(→Database entries) |
(→Basic information/ Evolution) |
||
| Line 70: | Line 70: | ||
* '''Protein family:''' RocA subfamily (according to Swiss-Prot) | * '''Protein family:''' RocA subfamily (according to Swiss-Prot) | ||
| − | * '''Paralogous protein(s):''' [[YcgN]] | + | * '''Paralogous protein(s):''' [[YcgN]], additional similar proteins: [[YfmT]], [[AldY]], [[YcbD]], [[GabD]], [[GbsA]], [[DhaS]], [[IolA]], [[AldX]], [[YwdH]] |
=== Extended information on the protein === | === Extended information on the protein === | ||
Revision as of 22:21, 29 June 2011
- Description: 3-hydroxy-1-pyrroline-5-carboxylate dehydrogenase
| Gene name | rocA |
| Synonyms | ipa-76d |
| Essential | no |
| Product | 3-hydroxy-1-pyrroline-5-carboxylate dehydrogenase |
| Function | arginine, ornithine and citrulline utilization |
| Metabolic function and regulation of this protein in SubtiPathways: Ammonium/ glutamate | |
| MW, pI | 56 kDa, 5.58 |
| Gene length, protein length | 1545 bp, 515 aa |
| Immediate neighbours | rocB, rocG |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 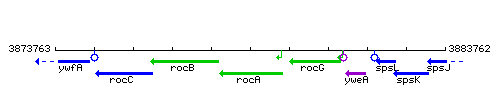
This image was kindly provided by SubtiList
| |
Contents
Categories containing this gene/protein
utilization of amino acids, phosphoproteins
This gene is a member of the following regulons
AhrC regulon, CodY regulon, RocR regulon, SigL regulon
The gene
Basic information
- Locus tag: BSU37780
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: (S)-1-pyrroline-5-carboxylate + NAD(P)+ + 2 H2O = L-glutamate + NAD(P)H (according to Swiss-Prot)
- Protein family: RocA subfamily (according to Swiss-Prot)
- Paralogous protein(s): YcgN, additional similar proteins: YfmT, AldY, YcbD, GabD, GbsA, DhaS, IolA, AldX, YwdH
Extended information on the protein
- Kinetic information:
- Domains:
- Modification: phosphorylation on (Thr-2 OR Thr-4 OR Tyr-5) PubMed
- Cofactor(s):
- Effectors of protein activity:
- Interactions:
- Localization:
Database entries
- Structure: 3RJL (1-pyrroline-5-carboxylate dehydrogenase from Bacillus licheniformis, 68% identity, 88% similarity)
- UniProt: P39634
- KEGG entry: [3]
- E.C. number: 1.5.1.12
Additional information
Expression and regulation
- Regulation:
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Additional publications: PubMed