Difference between revisions of "Sandbox"
| Line 1: | Line 1: | ||
| − | * '''Description:''' | + | * '''Description:''' aconitate hydratase (aconitase)<br/><br/> |
{| align="right" border="1" cellpadding="2" | {| align="right" border="1" cellpadding="2" | ||
|- | |- | ||
|style="background:#ABCDEF;" align="center"|'''Gene name''' | |style="background:#ABCDEF;" align="center"|'''Gene name''' | ||
| − | |'' | + | |''citB'' |
|- | |- | ||
|style="background:#ABCDEF;" align="center"| '''Synonyms''' || '' '' | |style="background:#ABCDEF;" align="center"| '''Synonyms''' || '' '' | ||
|- | |- | ||
| − | |style="background:#ABCDEF;" align="center"| '''Essential''' || no | + | |style="background:#ABCDEF;" align="center"| '''Essential''' || no |
|- | |- | ||
| − | |style="background:#ABCDEF;" align="center"| '''Product''' || | + | |style="background:#ABCDEF;" align="center"| '''Product''' || aconitate hydratase (aconitase) |
|- | |- | ||
| − | |style="background:#ABCDEF;" align="center"|'''Function''' || | + | |style="background:#ABCDEF;" align="center"|'''Function''' || TCA cycle |
|- | |- | ||
| − | |style="background:#ABCDEF;" align="center"| '''MW, pI''' || | + | |style="background:#ABCDEF;" align="center"| '''MW, pI''' || 99 kDa, 4.903 |
|- | |- | ||
| − | |style="background:#ABCDEF;" align="center"| '''Gene length, protein length''' || | + | |style="background:#ABCDEF;" align="center"| '''Gene length, protein length''' || 2727 bp, 909 aa |
|- | |- | ||
| − | |style="background:#ABCDEF;" align="center"|'''Immediate neighbours''' || ''[[ | + | |style="background:#ABCDEF;" align="center"|'''Immediate neighbours''' || ''[[sspO]], [[yneN]]'' |
|- | |- | ||
|colspan="2" style="background:#FAF8CC;" align="center"|'''Hier soll was neues rein''' | |colspan="2" style="background:#FAF8CC;" align="center"|'''Hier soll was neues rein''' | ||
|- | |- | ||
| − | |colspan="2" | '''Genetic context''' <br/> [[Image: | + | |colspan="2" | '''Genetic context''' <br/> [[Image:citB_context.gif]] |
<div align="right"> <small>This image was kindly provided by [http://genolist.pasteur.fr/SubtiList/ SubtiList]</small></div> | <div align="right"> <small>This image was kindly provided by [http://genolist.pasteur.fr/SubtiList/ SubtiList]</small></div> | ||
|- | |- | ||
| Line 38: | Line 38: | ||
===Phenotypes of a mutant === | ===Phenotypes of a mutant === | ||
| + | glutamate auxotrophy and a defect in sporulation [http://www.ncbi.nlm.nih.gov/pubmed/9393699 PubMed] | ||
=== Database entries === | === Database entries === | ||
| − | * '''DBTBS entry:''' [http://dbtbs.hgc.jp/COG/prom/ | + | * '''DBTBS entry:''' [http://dbtbs.hgc.jp/COG/prom/citB.html] |
| − | * '''SubtiList entry:''' [http://genolist.pasteur.fr/SubtiList/genome.cgi?gene_detail+ | + | * '''SubtiList entry:''' [http://genolist.pasteur.fr/SubtiList/genome.cgi?gene_detail+BG10478] |
=== Additional information=== | === Additional information=== | ||
| Line 52: | Line 53: | ||
=== Basic information/ Evolution === | === Basic information/ Evolution === | ||
| − | * '''Catalyzed reaction/ biological activity:''' | + | * '''Catalyzed reaction/ biological activity:''' : Citrate <---> Isocitrate, binding to iron responsive elements (IRE RNA) in the absence of the FeS cluster [http://www.ncbi.nlm.nih.gov/pubmed/10468622 PubMed] |
* '''Protein family:''' | * '''Protein family:''' | ||
| Line 66: | Line 67: | ||
* '''Modification:''' | * '''Modification:''' | ||
| − | * '''Cofactor(s):''' | + | * '''Cofactor(s):''' FeS cluster |
* '''Effectors of protein activity:''' | * '''Effectors of protein activity:''' | ||
| Line 76: | Line 77: | ||
=== Database entries === | === Database entries === | ||
| − | * '''Structure:''' | + | * '''Structure:''' [http://www.rcsb.org/pdb/cgi/explore.cgi?pdbId=1L5J 1L5J] (''E. coli'') |
* '''Swiss prot entry:''' | * '''Swiss prot entry:''' | ||
| − | * '''KEGG entry:''' [http://www.genome.jp/dbget-bin/www_bget?bsu+ | + | * '''KEGG entry:''' [http://www.genome.jp/dbget-bin/www_bget?bsu+BSU18000] |
| − | * '''E.C. number:''' | + | * '''E.C. number:''' 4.2.1.3 |
=== Additional information=== | === Additional information=== | ||
| + | ''B. subtilis'' aconitase is both an enzyme and an RNA binding protein [http://www.ncbi.nlm.nih.gov/pubmed/10468622 PubMed] | ||
=Expression and regulation= | =Expression and regulation= | ||
| − | * '''Operon:''' | + | * '''Operon:''' ''[[citB]]'' |
| − | * ''' | + | * '''Sigma factor:''' [[SigA]] |
| − | * '''Regulation:''' | + | * '''Regulation:''' repressed by glucose (3.7-fold) ([[CcpA]]) [http://www.ncbi.nlm.nih.gov/pubmed/12850135 PubMed], repression by glucose + arginine ([[CcpC]]) [http://www.ncbi.nlm.nih.gov/sites/entrez/16395550 PubMed] |
| − | * '''Regulatory mechanism:''' | + | * '''Regulatory mechanism:''' [[CcpA]]: transcription repression, [[CcpC]]: transcription repression [http://www.ncbi.nlm.nih.gov/sites/entrez/12100558 PubMed], [[AbrB]]: transcription activation [http://www.ncbi.nlm.nih.gov/pubmed/12591885 PubMed] |
| − | * '''Additional information:''' | + | * '''Additional information:''' |
=Biological materials = | =Biological materials = | ||
| − | * '''Mutant:''' | + | * '''Mutant:''' GP683 (erm), available in [[Stülke]] lab |
* '''Expression vector:''' | * '''Expression vector:''' | ||
| Line 113: | Line 115: | ||
=Labs working on this gene/protein= | =Labs working on this gene/protein= | ||
| + | |||
| + | [[Linc Sonenshein|Linc Sonenshein]], Tufts University, Boston, MA, USA [http://www.tufts.edu/sackler/microbiology/faculty/sonenshein/index.html Homepage] | ||
| + | |||
| + | [[Stülke|Jörg Stülke]], University of Göttingen, Germany | ||
| + | [http://wwwuser.gwdg.de/~genmibio/stuelke.html Homepage] | ||
=Your additional remarks= | =Your additional remarks= | ||
| Line 118: | Line 125: | ||
=References= | =References= | ||
| + | # Blencke et al. (2003) Transcriptional profiling of gene expression in response to glucose in ''Bacillus subtilis'': regulation of the central metabolic pathways. ''Metab Eng.'' '''5:''' 133-149 [http://www.ncbi.nlm.nih.gov/pubmed/12850135 PubMed] | ||
| + | # Rosenkrantz MS, Dingman DW, Sonenshein AL (1985) Bacillus subtilis citB gene is regulated synergistically by glucose and glutamine. J Bacteriol 164:155-164. [http://www.ncbi.nlm.nih.gov/sites/entrez/2413006 PubMed] | ||
| + | # Jourlin-Castelli C, Mani N, Nakano MM, Sonenshein AL (2000) CcpC, a novel regulator of the LysR family required for glucose repression of the citB gene in Bacillus subtilis. J Mol Biol 295:865-878. [http://www.ncbi.nlm.nih.gov/sites/entrez/10656796 PubMed] | ||
| + | # Alén C, Sonenshein AL (1999) Bacillus subtilis aconitase is an RNA-binding protein. Proc Natl Acad Sci USA 96:10412-10417. [http://www.ncbi.nlm.nih.gov/sites/entrez/10468622 PubMed] | ||
| + | # Fisher SH, Magasanik B (1984) 2-Ketoglutarate and the regulation of aconitase and histidase formation Bacillus subtilis. J Bacteriol 158:379-382. [http://www.ncbi.nlm.nih.gov/sites/entrez/6143742 PubMed] | ||
| + | # Kim, H. J., S. I. Kim, M. Ratnayake-Lecamwasam, K. Tachikawa, A. L. Sonenshein, and M. Strauch. (2003) Complex regulation of the Bacillus subtilis aconitase gene. J. Bacteriol. 185:1672-1680. [http://www.ncbi.nlm.nih.gov/sites/entrez/12591885 PubMed] | ||
| + | # Blencke, H.-M., Reif, I., Commichau, F. M., Detsch, C., Wacker, I., Ludwig, H. & Stülke, J. (2006) Regulation of citB expression in Bacillus subtilis: Integration of multiple metabolic signals in the citrate pool and by the general nitrogen regulatory system. Arch. Microbiol. 185: 136-146. [http://www.ncbi.nlm.nih.gov/sites/entrez/16395550 PubMed] | ||
| + | # Serio, A. W., Pechter, K. B., and Sonenshein, A. L. (2006) Bacillus subtilis aconitase is required for efficient late-sporulation gene expression. J Bacteriol 188: 6396-6405. [http://www.ncbi.nlm.nih.gov/sites/entrez/16923907 PubMed] | ||
| + | # Nakano MM, Zuber P, Sonenshein AL (1998) Anaerobic regulation of ''Bacillus subtilis'' Krebs cycle genes. J Bacteriol. 180:3304-3311. [http://www.ncbi.nlm.nih.gov/pubmed/9642180 PubMed] | ||
| + | # Craig JE, Ford MJ, Blaydon DC, Sonenshein AL (1997) A null mutation in the ''Bacillus subtilis'' aconitase gene causes a block in Spo0A-phosphate-dependent gene expression. J Bacteriol 197:7351-7359. [http://www.ncbi.nlm.nih.gov/pubmed/9393699 PubMed] | ||
| + | # Kim HJ, Kim SI, Ratnayake-Lecamwasam M, Tachikawa K, Sonenshein AL, Strauch M (2003) Complex regulation of the ''Bacillus subtilis'' aconitase gene. J Bacteriol. 185:1672-1680. [http://www.ncbi.nlm.nih.gov/pubmed/12591885 PubMed] | ||
| + | # Fouet, A., and Sonenshein, A. L. (1990) A target for carbon source-dependent negative regulation of the ''citB'' promoter of ''Bacillus subtilis''. J Bacteriol 172: 835-844. [http://www.ncbi.nlm.nih.gov/sites/entrez/2105305 PubMed] | ||
# Author1, Author2 & Author3 (year) Title ''Journal'' '''volume:''' page-page. [http://www.ncbi.nlm.nih.gov/sites/entrez/PMID PubMed] | # Author1, Author2 & Author3 (year) Title ''Journal'' '''volume:''' page-page. [http://www.ncbi.nlm.nih.gov/sites/entrez/PMID PubMed] | ||
Revision as of 18:38, 15 April 2009
- Description: aconitate hydratase (aconitase)
| Gene name | citB |
| Synonyms | |
| Essential | no |
| Product | aconitate hydratase (aconitase) |
| Function | TCA cycle |
| MW, pI | 99 kDa, 4.903 |
| Gene length, protein length | 2727 bp, 909 aa |
| Immediate neighbours | sspO, yneN |
| Hier soll was neues rein | |
Genetic context 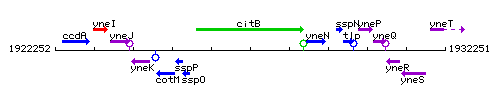
This image was kindly provided by SubtiList
| |
Contents
The gene
Basic information
- Coordinates:
Phenotypes of a mutant
glutamate auxotrophy and a defect in sporulation PubMed
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: : Citrate <---> Isocitrate, binding to iron responsive elements (IRE RNA) in the absence of the FeS cluster PubMed
- Protein family:
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s): FeS cluster
- Effectors of protein activity:
- Interactions:
- Localization:
Database entries
- Structure: 1L5J (E. coli)
- Swiss prot entry:
- KEGG entry: [3]
- E.C. number: 4.2.1.3
Additional information
B. subtilis aconitase is both an enzyme and an RNA binding protein PubMed
Expression and regulation
- Operon: citB
- Sigma factor: SigA
- Regulation: repressed by glucose (3.7-fold) (CcpA) PubMed, repression by glucose + arginine (CcpC) PubMed
- Regulatory mechanism: CcpA: transcription repression, CcpC: transcription repression PubMed, AbrB: transcription activation PubMed
- Additional information:
Biological materials
- Mutant: GP683 (erm), available in Stülke lab
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Linc Sonenshein, Tufts University, Boston, MA, USA Homepage
Jörg Stülke, University of Göttingen, Germany Homepage
Your additional remarks
References
- Blencke et al. (2003) Transcriptional profiling of gene expression in response to glucose in Bacillus subtilis: regulation of the central metabolic pathways. Metab Eng. 5: 133-149 PubMed
- Rosenkrantz MS, Dingman DW, Sonenshein AL (1985) Bacillus subtilis citB gene is regulated synergistically by glucose and glutamine. J Bacteriol 164:155-164. PubMed
- Jourlin-Castelli C, Mani N, Nakano MM, Sonenshein AL (2000) CcpC, a novel regulator of the LysR family required for glucose repression of the citB gene in Bacillus subtilis. J Mol Biol 295:865-878. PubMed
- Alén C, Sonenshein AL (1999) Bacillus subtilis aconitase is an RNA-binding protein. Proc Natl Acad Sci USA 96:10412-10417. PubMed
- Fisher SH, Magasanik B (1984) 2-Ketoglutarate and the regulation of aconitase and histidase formation Bacillus subtilis. J Bacteriol 158:379-382. PubMed
- Kim, H. J., S. I. Kim, M. Ratnayake-Lecamwasam, K. Tachikawa, A. L. Sonenshein, and M. Strauch. (2003) Complex regulation of the Bacillus subtilis aconitase gene. J. Bacteriol. 185:1672-1680. PubMed
- Blencke, H.-M., Reif, I., Commichau, F. M., Detsch, C., Wacker, I., Ludwig, H. & Stülke, J. (2006) Regulation of citB expression in Bacillus subtilis: Integration of multiple metabolic signals in the citrate pool and by the general nitrogen regulatory system. Arch. Microbiol. 185: 136-146. PubMed
- Serio, A. W., Pechter, K. B., and Sonenshein, A. L. (2006) Bacillus subtilis aconitase is required for efficient late-sporulation gene expression. J Bacteriol 188: 6396-6405. PubMed
- Nakano MM, Zuber P, Sonenshein AL (1998) Anaerobic regulation of Bacillus subtilis Krebs cycle genes. J Bacteriol. 180:3304-3311. PubMed
- Craig JE, Ford MJ, Blaydon DC, Sonenshein AL (1997) A null mutation in the Bacillus subtilis aconitase gene causes a block in Spo0A-phosphate-dependent gene expression. J Bacteriol 197:7351-7359. PubMed
- Kim HJ, Kim SI, Ratnayake-Lecamwasam M, Tachikawa K, Sonenshein AL, Strauch M (2003) Complex regulation of the Bacillus subtilis aconitase gene. J Bacteriol. 185:1672-1680. PubMed
- Fouet, A., and Sonenshein, A. L. (1990) A target for carbon source-dependent negative regulation of the citB promoter of Bacillus subtilis. J Bacteriol 172: 835-844. PubMed
- Author1, Author2 & Author3 (year) Title Journal volume: page-page. PubMed