Difference between revisions of "MtnA"
| Line 14: | Line 14: | ||
|style="background:#ABCDEF;" align="center"|'''Function''' || methionine salvage | |style="background:#ABCDEF;" align="center"|'''Function''' || methionine salvage | ||
|- | |- | ||
| − | |colspan="2" style="background:#FAF8CC;" align="center"| '''Gene expression levels in [http:// | + | |colspan="2" style="background:#FAF8CC;" align="center"| '''Gene expression levels in [http://subtiwiki.uni-goettingen.de/apps/expression/ ''Subti''Express]''': [http://subtiwiki.uni-goettingen.de/apps/expression/expression.php?search=BSU13550 mtnA] |
|- | |- | ||
|colspan="2" style="background:#FAF8CC;" align="center"| '''Metabolic function and regulation of this protein in [[SubtiPathways|''Subti''Pathways]]: <br/>[http://subtiwiki.uni-goettingen.de/pathways/cys_meth_and_sulfate_assimilation.html Cys, Met & Sulfate assimilation]''' | |colspan="2" style="background:#FAF8CC;" align="center"| '''Metabolic function and regulation of this protein in [[SubtiPathways|''Subti''Pathways]]: <br/>[http://subtiwiki.uni-goettingen.de/pathways/cys_meth_and_sulfate_assimilation.html Cys, Met & Sulfate assimilation]''' | ||
| Line 24: | Line 24: | ||
|style="background:#ABCDEF;" align="center"|'''Immediate neighbours''' || ''[[ogt]]'', ''[[mtnK]]'' | |style="background:#ABCDEF;" align="center"|'''Immediate neighbours''' || ''[[ogt]]'', ''[[mtnK]]'' | ||
|- | |- | ||
| − | | | + | |style="background:#FAF8CC;" align="center"|'''Sequences'''||[http://bsubcyc.org/BSUB/sequence-aa?type=GENE&object=BSU13550 Protein] [http://bsubcyc.org/BSUB/sequence?type=GENE&object=BSU13550 DNA] [http://bsubcyc.org/BSUB/seq-selector?chromosome=CHROM-1&object=BSU13550 Advanced_DNA] |
|- | |- | ||
|colspan="2" | '''Genetic context''' <br/> [[Image:ykrS_context.gif]] | |colspan="2" | '''Genetic context''' <br/> [[Image:ykrS_context.gif]] | ||
Revision as of 13:39, 13 May 2013
- Description: 5-methylthioribose-1-phosphate isomerase
| Gene name | mtnA |
| Synonyms | ykrS, mtnS |
| Essential | no |
| Product | 5-methylthioribose-1-phosphate isomerase |
| Function | methionine salvage |
| Gene expression levels in SubtiExpress: mtnA | |
| Metabolic function and regulation of this protein in SubtiPathways: Cys, Met & Sulfate assimilation | |
| MW, pI | 38 kDa, 5.003 |
| Gene length, protein length | 1059 bp, 353 aa |
| Immediate neighbours | ogt, mtnK |
| Sequences | Protein DNA Advanced_DNA |
Genetic context 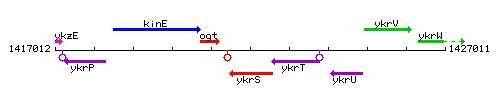
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed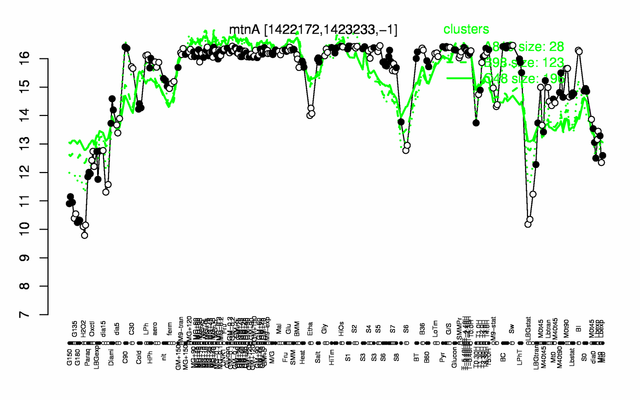
| |
Contents
Categories containing this gene/protein
biosynthesis/ acquisition of amino acids, phosphoproteins
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU13550
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: S-methyl-5-thio-alpha-D-ribose 1-phosphate = S-methyl-5-thio-D-ribulose 1-phosphate (according to Swiss-Prot)
- Protein family: MtnA subfamily (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- phosphorylated on Arg-80 and Arg-298 PubMed
- Cofactor(s):
- Effectors of protein activity:
Database entries
- Structure: 2YVK
- UniProt: O31662
- KEGG entry: [3]
- E.C. number: 5.3.1.23
Additional information
- subject to Clp-dependent proteolysis upon glucose starvation PubMed
Expression and regulation
- Regulatory mechanism: S-box: transcription termination/ antitermination, the S-box riboswitch binds S-adenosylmethionine resulting in termination PubMed
- Additional information: subject to Clp-dependent proteolysis upon glucose starvation PubMed
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Alexander K W Elsholz, Kürsad Turgay, Stephan Michalik, Bernd Hessling, Katrin Gronau, Dan Oertel, Ulrike Mäder, Jörg Bernhardt, Dörte Becher, Michael Hecker, Ulf Gerth
Global impact of protein arginine phosphorylation on the physiology of Bacillus subtilis.
Proc Natl Acad Sci U S A: 2012, 109(19);7451-6
[PubMed:22517742]
[WorldCat.org]
[DOI]
(I p)
Jerneja Tomsic, Brooke A McDaniel, Frank J Grundy, Tina M Henkin
Natural variability in S-adenosylmethionine (SAM)-dependent riboswitches: S-box elements in bacillus subtilis exhibit differential sensitivity to SAM In vivo and in vitro.
J Bacteriol: 2008, 190(3);823-33
[PubMed:18039762]
[WorldCat.org]
[DOI]
(I p)
Agnieszka Sekowska, Valérie Dénervaud, Hiroki Ashida, Karine Michoud, Dieter Haas, Akiho Yokota, Antoine Danchin
Bacterial variations on the methionine salvage pathway.
BMC Microbiol: 2004, 4;9
[PubMed:15102328]
[WorldCat.org]
[DOI]
(I e)
Maumita Mandal, Benjamin Boese, Jeffrey E Barrick, Wade C Winkler, Ronald R Breaker
Riboswitches control fundamental biochemical pathways in Bacillus subtilis and other bacteria.
Cell: 2003, 113(5);577-86
[PubMed:12787499]
[WorldCat.org]
[DOI]
(P p)
Ulrike Mäder, Georg Homuth, Christian Scharf, Knut Büttner, Rüdiger Bode, Michael Hecker
Transcriptome and proteome analysis of Bacillus subtilis gene expression modulated by amino acid availability.
J Bacteriol: 2002, 184(15);4288-95
[PubMed:12107147]
[WorldCat.org]
[DOI]
(P p)
Brooke A Murphy, Frank J Grundy, Tina M Henkin
Prediction of gene function in methylthioadenosine recycling from regulatory signals.
J Bacteriol: 2002, 184(8);2314-8
[PubMed:11914366]
[WorldCat.org]
[DOI]
(P p)
A Sekowska, L Mulard, S Krogh, J K Tse, A Danchin
MtnK, methylthioribose kinase, is a starvation-induced protein in Bacillus subtilis.
BMC Microbiol: 2001, 1;15
[PubMed:11545674]
[WorldCat.org]
[DOI]
(I p)