Difference between revisions of "Spx"
(→Categories containing this gene/protein) |
|||
| Line 136: | Line 136: | ||
** post-translational control by [[ClpX]]-[[ClpP]]: Spx naturally contains a C-terminal sequence that resembles the [[SsrA]] tag and targets the protein for degradation. [http://www.ncbi.nlm.nih.gov/pubmed/12642660?ordinalpos=27&itool=EntrezSystem2.PEntrez.Pubmed.Pubmed_ResultsPanel.Pubmed_DefaultReportPanel.Pubmed_RVDocSum PubMed] | ** post-translational control by [[ClpX]]-[[ClpP]]: Spx naturally contains a C-terminal sequence that resembles the [[SsrA]] tag and targets the protein for degradation. [http://www.ncbi.nlm.nih.gov/pubmed/12642660?ordinalpos=27&itool=EntrezSystem2.PEntrez.Pubmed.Pubmed_ResultsPanel.Pubmed_DefaultReportPanel.Pubmed_RVDocSum PubMed] | ||
** proteolysis is enhanced by [[YjbH]] [http://www.ncbi.nlm.nih.gov/pubmed/19074380?ordinalpos=1&itool=EntrezSystem2.PEntrez.Pubmed.Pubmed_ResultsPanel.Pubmed_DefaultReportPanel.Pubmed_RVDocSum PubMed] and counter-acted by [[YirB]] {{PubMed|21378193}} | ** proteolysis is enhanced by [[YjbH]] [http://www.ncbi.nlm.nih.gov/pubmed/19074380?ordinalpos=1&itool=EntrezSystem2.PEntrez.Pubmed.Pubmed_ResultsPanel.Pubmed_DefaultReportPanel.Pubmed_RVDocSum PubMed] and counter-acted by [[YirB]] {{PubMed|21378193}} | ||
| + | ** the mRNA is substantially stabilized upon depletion of [[Rny|RNase Y]] {{PubMed|21815947}} | ||
=Biological materials = | =Biological materials = | ||
| Line 170: | Line 171: | ||
==Original Publications== | ==Original Publications== | ||
'''Additional publications:''' {{PubMed|21378193}} | '''Additional publications:''' {{PubMed|21378193}} | ||
| + | <big>''Lehnik-Habrink M, Schaffer M, Mäder U, Diethmaier C, Herzberg C, Stülke J'' </big> | ||
| + | <big>'''RNA processing in ''Bacillus subtilis'': identification of targets of the essential RNase Y.''' </big> | ||
| + | <big>Mol Microbiol. 2011 81(6): 1459-1473. </big> | ||
| + | [http://www.ncbi.nlm.nih.gov/pubmed/21815947 PubMed:21815947] | ||
<pubmed> 18487332,17908206,17434969,17158660, 19074380, 15805528, 18662407, 16885442, 18179421, 17434969, 17908206, 17158660, 11703662, 15659166, 12642660, 12057962, 10482513, 18687074, 12775685, 16740936, 17827297, 20084284 20057163 10913081 </pubmed> | <pubmed> 18487332,17908206,17434969,17158660, 19074380, 15805528, 18662407, 16885442, 18179421, 17434969, 17908206, 17158660, 11703662, 15659166, 12642660, 12057962, 10482513, 18687074, 12775685, 16740936, 17827297, 20084284 20057163 10913081 </pubmed> | ||
[[Category:Protein-coding genes]] | [[Category:Protein-coding genes]] | ||
Revision as of 20:21, 19 November 2011
- Description: transcriptional regulator Spx, involved in regulation of many genes.
| Gene name | spx |
| Synonyms | yjbD |
| Essential | no |
| Product | transcriptional regulator Spx |
| Function | negative and positive regulator of many genes |
| Interactions involving this protein in SubtInteract: Spx | |
| Metabolic function and regulation of this protein in SubtiPathways: Riboflavin / FAD | |
| MW, pI | 15,5 kDa, 7.80 |
| Gene length, protein length | 393 bp, 131 amino acids |
| Immediate neighbours | yjbC, yjbE |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 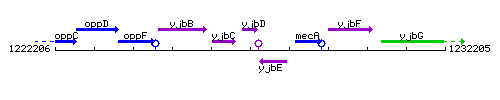
This image was kindly provided by SubtiList
| |
Contents
Categories containing this gene/protein
transcription factors and their control, general stress proteins (controlled by SigB), cell envelope stress proteins (controlled by SigM, V, W, X, Y)
This gene is a member of the following regulons
PerR regulon, SigB regulon, SigM regulon, SigW regulon, SigX regulon
The Spx regulon
The gene
Basic information
- Locus tag: BSU11500
Phenotypes of a mutant
Loss of up-regulation of the methionine sulfoxide reductase (msrA-msrB) operon in response to thiol specific oxidative stress, also loss of trxA and trxB upregulation in response to thiol specific oxidative stress.
Database entries
- DBTBS entry: [1]
- SubtiList entry: link
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- transcriptional regulator of many genes in response to thiol specific oxidative stress (transcription activator of trxA and trxB)
- in addition, Spx inhibits transcription by binding to the C-terminal domain of the alpha subunit of RNAP (RpoA), disrupting complex formation between RNAP and certain transcriptional activator proteins like ResD and ComA
- in response to thiol specific oxidative stress, Spx can also activate transcription, making it a general regulator that exerts both positive and negative control over transcription initiation
- involved in competence regulation PubMed
- Protein family: Spx subfamily (according to Swiss-Prot) Arsenate Reductase (ArsC) family, Spx subfamily
- Paralogous protein(s): MgsR
Extended information on the protein
- Kinetic information:
- Domains: CXXC (10-13): Acts as a disulfide switch for the redox-sensitive transcriptional regulation of genes that function in thiol homeostasis.
- Modification: Cysteine oxidation of the CXXC motif
- Cofactor(s):
- Effectors of protein activity:
- Localization: cytoplasm (according to Swiss-Prot)
Database entries
- UniProt: O31602
- KEGG entry: [2]
- E.C. number:
Additional information
Expression and regulation
- Regulation:
- Additional information:
- post-translational control by ClpX-ClpP: Spx naturally contains a C-terminal sequence that resembles the SsrA tag and targets the protein for degradation. PubMed
- proteolysis is enhanced by YjbH PubMed and counter-acted by YirB PubMed
- the mRNA is substantially stabilized upon depletion of RNase Y PubMed
Biological materials
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system: B. pertussis adenylate cyclase-based bacterial two hybrid system (BACTH), available in Stülke lab
- Antibody:
Labs working on this gene/protein
Peter Zuber, Oregon Health and Science University, USA Homepage
Richard Brennan, Houston, Texas, USA Homepage
Your additional remarks
References
Reviews
Additional reviews: PubMed
The Spx regulon
Structural analysis of Spx
Original Publications
Additional publications: PubMed
Lehnik-Habrink M, Schaffer M, Mäder U, Diethmaier C, Herzberg C, Stülke J RNA processing in Bacillus subtilis: identification of targets of the essential RNase Y. Mol Microbiol. 2011 81(6): 1459-1473. PubMed:21815947