Difference between revisions of "SigX"
(→Other original publications) |
|||
| Line 137: | Line 137: | ||
'''Additional publications:''' {{PubMed|20817771}} | '''Additional publications:''' {{PubMed|20817771}} | ||
==Other original publications== | ==Other original publications== | ||
| − | <pubmed>11222589,9139908,14993308,12399481,17675383,9683469,14762009,19047346,14651641,10960106,,19465659 , 9636707 , 16306698 7934830 19767430 9393439 | + | <pubmed>11222589,9139908,14993308,12399481,17675383,9683469,14762009,19047346,14651641,10960106,,19465659 , 9636707 , 16306698 7934830 19767430 9393439 </pubmed> |
[[Category:Protein-coding genes]] | [[Category:Protein-coding genes]] | ||
Revision as of 21:13, 10 December 2010
- Description: RNA polymerase ECF-type sigma factor SigX
| Gene name | sigX |
| Synonyms | ypuM |
| Essential | no |
| Product | RNA polymerase ECF-type sigma factor SigX |
| Function | cell surface properties |
| MW, pI | 23 kDa, 6.086 |
| Gene length, protein length | 582 bp, 194 aa |
| Immediate neighbours | rsiX, resE |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 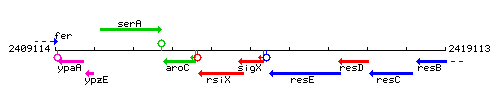
This image was kindly provided by SubtiList
| |
Contents
Categories containing this gene/protein
transcription, sigma factors and their control, cell envelope stress proteins (controlled by SigM, W, X, Y), membrane proteins
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU23100
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family: ECF subfamily (according to Swiss-Prot)
- Paralogous protein(s):
Genes controlled by SigX
abh, pbpX, csbB, abh, dltA-dltB-dltC-dltD-dltE
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Localization: cell membrane (according to Swiss-Prot)
Database entries
- Structure:
- UniProt: P35165
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Regulation:
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Additional publications: PubMed
Other original publications