Difference between revisions of "RplC"
| Line 95: | Line 95: | ||
* '''[[Localization]]:''' | * '''[[Localization]]:''' | ||
| + | ** located at the peptidyl transferase center of the 50S ribosomal subunit {{PubMed|23759864}} | ||
=== Database entries === | === Database entries === | ||
| Line 142: | Line 143: | ||
=References= | =References= | ||
| − | + | <pubmed>17981968,9371452,19154332, 19653700 11948165, 8635744 23175651 23759864 23002217</pubmed> | |
| − | <pubmed>17981968,9371452,19154332, 19653700 11948165, 8635744 23175651</pubmed> | ||
[[Category:Protein-coding genes]] | [[Category:Protein-coding genes]] | ||
Revision as of 16:05, 14 June 2013
- Description: ribosomal protein L3
| Gene name | rplC |
| Synonyms | |
| Essential | yes PubMed |
| Product | ribosomal protein L3 (BL3) |
| Function | translation |
| Gene expression levels in SubtiExpress: rplC | |
| Interactions involving this protein in SubtInteract: RplC | |
| MW, pI | 22 kDa, 10.31 |
| Gene length, protein length | 627 bp, 209 aa |
| Immediate neighbours | rpsJ, rplD |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 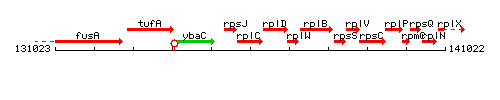
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed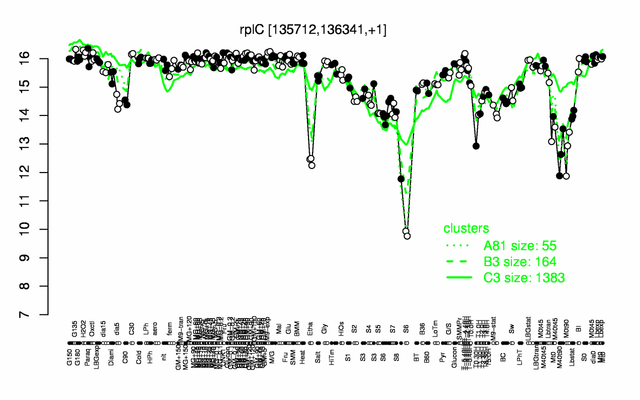
| |
Contents
Categories containing this gene/protein
translation, essential genes, universally conserved proteins
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU01160
Phenotypes of a mutant
essential PubMed
Database entries
- DBTBS entry: no entry
- SubtiList entry: [1]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: RplC binding to the precursor 23S rRNA stimulates MrnC activity PubMed
- Protein family: ribosomal protein L3P family (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Localization:
- located at the peptidyl transferase center of the 50S ribosomal subunit PubMed
Database entries
- Structure:
- UniProt: P42920
- KEGG entry: [2]
- E.C. number:
Additional information
Expression and regulation
- Operon: rpsJ-rplC-rplD-rplW-rplB-rpsS-rplV-rpsC-rplP-rpmC-rpsQ-rplN-rplX-rplE-rpsN-rpsH-rplF-rplR-rpsE-rpmD-rplO PubMed
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system: B. pertussis adenylate cyclase-based bacterial two hybrid system (BACTH), available in Stülke lab
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Genki Akanuma, Shota Suzuki, Koichi Yano, Hideaki Nanamiya, Yousuke Natori, Eri Namba, Kazuya Watanabe, Kazumi Tagami, Takuya Takeda, Yuka Iizuka, Ako Kobayashi, Morio Ishizuka, Hirofumi Yoshikawa, Fujio Kawamura
Single mutations introduced in the essential ribosomal proteins L3 and S10 cause a sporulation defect in Bacillus subtilis.
J Gen Appl Microbiol: 2013, 59(2);105-17
[PubMed:23759864]
[WorldCat.org]
[DOI]
(I p)
Martin Lehnik-Habrink, Leonie Rempeters, Ákos T Kovács, Christoph Wrede, Claudia Baierlein, Heike Krebber, Oscar P Kuipers, Jörg Stülke
DEAD-Box RNA helicases in Bacillus subtilis have multiple functions and act independently from each other.
J Bacteriol: 2013, 195(3);534-44
[PubMed:23175651]
[WorldCat.org]
[DOI]
(I p)
Genki Akanuma, Hideaki Nanamiya, Yousuke Natori, Koichi Yano, Shota Suzuki, Shuya Omata, Morio Ishizuka, Yasuhiko Sekine, Fujio Kawamura
Inactivation of ribosomal protein genes in Bacillus subtilis reveals importance of each ribosomal protein for cell proliferation and cell differentiation.
J Bacteriol: 2012, 194(22);6282-91
[PubMed:23002217]
[WorldCat.org]
[DOI]
(I p)
Matthew A Lauber, William E Running, James P Reilly
B. subtilis ribosomal proteins: structural homology and post-translational modifications.
J Proteome Res: 2009, 8(9);4193-206
[PubMed:19653700]
[WorldCat.org]
[DOI]
(P p)
Yulia Redko, Ciarán Condon
Ribosomal protein L3 bound to 23S precursor rRNA stimulates its maturation by Mini-III ribonuclease.
Mol Microbiol: 2009, 71(5);1145-54
[PubMed:19154332]
[WorldCat.org]
[DOI]
(I p)
Catherine Wicker-Planquart, Anne-Emmanuelle Foucher, Mathilde Louwagie, Robert A Britton, Jean-Michel Jault
Interactions of an essential Bacillus subtilis GTPase, YsxC, with ribosomes.
J Bacteriol: 2008, 190(2);681-90
[PubMed:17981968]
[WorldCat.org]
[DOI]
(I p)
Christine Eymann, Georg Homuth, Christian Scharf, Michael Hecker
Bacillus subtilis functional genomics: global characterization of the stringent response by proteome and transcriptome analysis.
J Bacteriol: 2002, 184(9);2500-20
[PubMed:11948165]
[WorldCat.org]
[DOI]
(P p)
X Li, L Lindahl, Y Sha, J M Zengel
Analysis of the Bacillus subtilis S10 ribosomal protein gene cluster identifies two promoters that may be responsible for transcription of the entire 15-kilobase S10-spc-alpha cluster.
J Bacteriol: 1997, 179(22);7046-54
[PubMed:9371452]
[WorldCat.org]
[DOI]
(P p)
J W Suh, S A Boylan, S H Oh, C W Price
Genetic and transcriptional organization of the Bacillus subtilis spc-alpha region.
Gene: 1996, 169(1);17-23
[PubMed:8635744]
[WorldCat.org]
[DOI]
(P p)