Difference between revisions of "SdhB"
| Line 13: | Line 13: | ||
|- | |- | ||
|style="background:#ABCDEF;" align="center"|'''Function''' || TCA cycle | |style="background:#ABCDEF;" align="center"|'''Function''' || TCA cycle | ||
| + | |- | ||
| + | |colspan="2" style="background:#FAF8CC;" align="center"| '''Gene expression levels in [http://cellpublisher.gobics.de/subtiexpress/ ''Subti''Express]''': [http://cellpublisher.gobics.de/subtiexpress/bsu/BSU28430 sdhB] | ||
|- | |- | ||
|colspan="2" style="background:#FAF8CC;" align="center"| '''Interactions involving this protein in [http://cellpublisher.gobics.de/subtinteract/startpage/start/ ''Subt''Interact]''': [http://cellpublisher.gobics.de/subtinteract/interactionList/2/SdhB SdhB] | |colspan="2" style="background:#FAF8CC;" align="center"| '''Interactions involving this protein in [http://cellpublisher.gobics.de/subtinteract/startpage/start/ ''Subt''Interact]''': [http://cellpublisher.gobics.de/subtinteract/interactionList/2/SdhB SdhB] | ||
Revision as of 15:37, 7 August 2012
- Description: succinate dehydrogenase
| Gene name | sdhB |
| Synonyms | |
| Essential | no |
| Product | succinate dehydrogenase (iron-sulfur protein) |
| Function | TCA cycle |
| Gene expression levels in SubtiExpress: sdhB | |
| Interactions involving this protein in SubtInteract: SdhB | |
| Metabolic function and regulation of this protein in SubtiPathways: Central C-metabolism | |
| MW, pI | 28 kDa, 7.989 |
| Gene length, protein length | 759 bp, 253 aa |
| Immediate neighbours | ysmA, sdhA |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 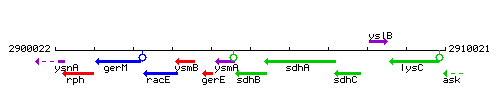
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed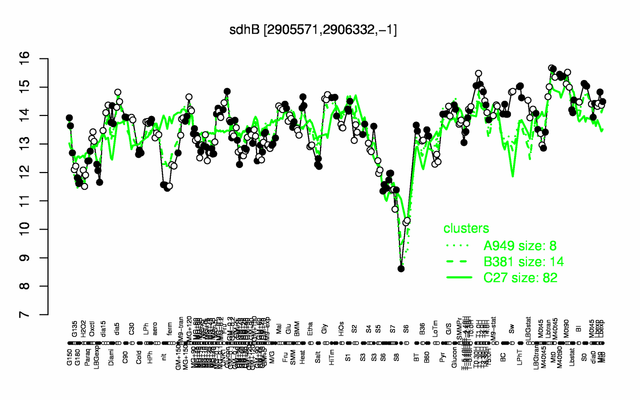
| |
Contents
Categories containing this gene/protein
carbon core metabolism, membrane proteins
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU28430
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: Succinate + acceptor = fumarate + reduced acceptor (according to Swiss-Prot)
- Protein family: succinate dehydrogenase/fumarate reductase iron-sulfur protein family (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s): Fe
- Effectors of protein activity:
- Localization: attached to the membrane PubMed
Database entries
- Structure: 1NEK (E. coli)
- UniProt: P08066
- KEGG entry: [3]
- E.C. number:EC 1.3.99.1
Additional information
- This enzyme is a trimer membrane-bound PubMed PubMed
- One subunit is bound to citochrome b558, and this subunit is the one bound to the cytosolic side of the membrane PubMed PubMed
- Another subunit is the flavoprotein one, required for FAD usage PubMed PubMed
- The other subunit has an iron-sulphur domain necessary for the catalytic activity PubMed PubMed
- extensive information on the structure and enzymatic properties of succinate dehydrogenase can be found at Proteopedia
Expression and regulation
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system: B. pertussis adenylate cyclase-based bacterial two hybrid system (BACTH), available in Stülke lab
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Gregory T Smaldone, Olga Revelles, Ahmed Gaballa, Uwe Sauer, Haike Antelmann, John D Helmann
A global investigation of the Bacillus subtilis iron-sparing response identifies major changes in metabolism.
J Bacteriol: 2012, 194(10);2594-605
[PubMed:22389480]
[WorldCat.org]
[DOI]
(I p)
Victoria Yankovskaya, Rob Horsefield, Susanna Törnroth, César Luna-Chavez, Hideto Miyoshi, Christophe Léger, Bernadette Byrne, Gary Cecchini, So Iwata
Architecture of succinate dehydrogenase and reactive oxygen species generation.
Science: 2003, 299(5607);700-4
[PubMed:12560550]
[WorldCat.org]
[DOI]
(I p)
C Hägerhäll, V Sled, L Hederstedt, T Ohnishi
The trinuclear iron-sulfur cluster S3 in Bacillus subtilis succinate:menaquinone reductase; effects of a mutation in the putative cluster ligation motif on enzyme activity and EPR properties.
Biochim Biophys Acta: 1995, 1229(3);356-62
[PubMed:7748886]
[WorldCat.org]
[DOI]
(P p)
C Hägerhäll, R Aasa, C von Wachenfeldt, L Hederstedt
Two hemes in Bacillus subtilis succinate:menaquinone oxidoreductase (complex II).
Biochemistry: 1992, 31(32);7411-21
[PubMed:1324713]
[WorldCat.org]
[DOI]
(P p)
L Melin, H Fridén, E Dehlin, L Rutberg, A von Gabain
The importance of the 5'-region in regulating the stability of sdh mRNA in Bacillus subtilis.
Mol Microbiol: 1990, 4(11);1881-9
[PubMed:1707123]
[WorldCat.org]
[DOI]
(P p)
L Melin, L Rutberg, A von Gabain
Transcriptional and posttranscriptional control of the Bacillus subtilis succinate dehydrogenase operon.
J Bacteriol: 1989, 171(4);2110-5
[PubMed:2495271]
[WorldCat.org]
[DOI]
(P p)
A AEvarsson, L Hederstedt
Ligands to the 2Fe iron-sulfur center in succinate dehydrogenase.
FEBS Lett: 1988, 232(2);298-302
[PubMed:2837411]
[WorldCat.org]
[DOI]
(P p)
L Melin, K Magnusson, L Rutberg
Identification of the promoter of the Bacillus subtilis sdh operon.
J Bacteriol: 1987, 169(7);3232-6
[PubMed:3036777]
[WorldCat.org]
[DOI]
(P p)
M K Phillips, L Hederstedt, S Hasnain, L Rutberg, J R Guest
Nucleotide sequence encoding the flavoprotein and iron-sulfur protein subunits of the Bacillus subtilis PY79 succinate dehydrogenase complex.
J Bacteriol: 1987, 169(2);864-73
[PubMed:3027051]
[WorldCat.org]
[DOI]
(P p)
S T Cole, C Condon, B D Lemire, J H Weiner
Molecular biology, biochemistry and bioenergetics of fumarate reductase, a complex membrane-bound iron-sulfur flavoenzyme of Escherichia coli.
Biochim Biophys Acta: 1985, 811(4);381-403
[PubMed:3910107]
[WorldCat.org]
[DOI]
(P p)
L Hederstedt, L Rutberg
Orientation of succinate dehydrogenase and cytochrome b558 in the Bacillus subtilis cytoplasmic membrane.
J Bacteriol: 1983, 153(1);57-65
[PubMed:6401289]
[WorldCat.org]
[DOI]
(P p)
L Hederstedt, L Rutberg
Succinate dehydrogenase--a comparative review.
Microbiol Rev: 1981, 45(4);542-55
[PubMed:6799760]
[WorldCat.org]
[DOI]
(P p)