DegU
- Description: two-component response regulator, regulation of degradative enzyme expression, genetic competence, biofilm formation, capsule biosynthesis (together with SwrAA), non-phosphorylated DegU is required for swarming motility
| Gene name | degU |
| Synonyms | sacU, iep |
| Essential | no |
| Product | two-component response regulator |
| Function | regulation of degradative enzymes, genetic competence,
biofilm formation and capsule synthesis |
| Gene expression levels in SubtiExpress: degU | |
| Interactions involving this protein in SubtInteract: DegU | |
| Metabolic function and regulation of this protein in SubtiPathways: DegU | |
| MW, pI | 25 kDa, 5.602 |
| Gene length, protein length | 687 bp, 229 aa |
| Immediate neighbours | yviA, degS |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 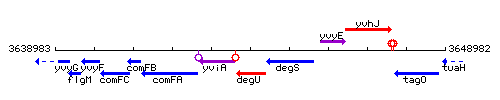
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed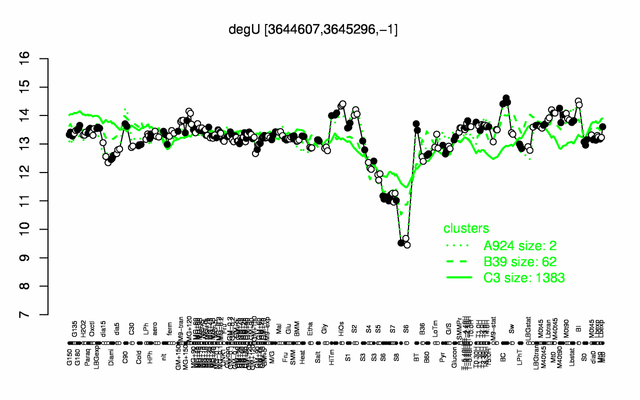
| |
Contents
Categories containing this gene/protein
biofilm formation, transcription factors and their control, phosphoproteins
This gene is a member of the following regulons
CcpA regulon, DegU regulon, SinR regulon, TnrA regulon
The DegU regulon
The gene
Basic information
- Locus tag: BSU35490
Phenotypes of a mutant
- defect in biofilm formation PubMed, this can be suppressed by DegU-independent expression of BslA PubMed
- the mutation suppresses the mucoid phenotype of motA or motB mutants due to reduced expression of the capB-capC-capA-capE operon PubMed
Database entries
- BsubCyc: BSU35490
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- transcription activator and repressor (see DegU regulon)
- DegU-P represses expression of the fla/che operon (flgB-flgC-fliE-fliF-fliG-fliH-fliI-fliJ-ylxF-fliK-flgD-flgE-fliL-fliM-fliY-cheY-fliZ-fliP-fliQ-fliR-flhB-flhA-flhF-flhG-cheB-cheA-cheW-cheC-cheD-sigD-swrB) in the absence of SwrA, and switches to an activator of the operon in the presence of SwrA PubMed
- Protein family:
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Modification:
- Effectors of protein activity:
- Localization: cytoplasm (according to Swiss-Prot)
Database entries
- BsubCyc: BSU35490
- Structure:
- UniProt: P13800
- KEGG entry: [3]
- E.C. number:
Additional information
- DegU-P is degraded by ClpC-ClpP PubMed
- accumulation of DegU-P results in decreased transcription of the epsA-epsO and tapA-sipW-tasA operons due to increased levels of Spo0A-P PubMed
Expression and regulation
- Regulatory mechanism:
- Additional information:
- DegU-P is degraded by ClpC-ClpP PubMed
- number of protein molecules per cell (minimal medium with glucose and ammonium): 1627 PubMed
- number of protein molecules per cell (complex medium with amino acids, without glucose): 2454 PubMed
- number of protein molecules per cell (minimal medium with glucose and ammonium, exponential phase): 2251 PubMed
- number of protein molecules per cell (minimal medium with glucose and ammonium, early stationary phase after glucose exhaustion): 1461 PubMed
- number of protein molecules per cell (minimal medium with glucose and ammonium, late stationary phase after glucose exhaustion): 1432 PubMed
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system: B. pertussis adenylate cyclase-based bacterial two hybrid system (BACTH), available in Jörg Stülke's lab
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Reviews
The DegU regulon
Original Publications
Fabian M Commichau, Achim Dickmanns, Jan Gundlach, Ralf Ficner, Jörg Stülke
A jack of all trades: the multiple roles of the unique essential second messenger cyclic di-AMP.
Mol Microbiol: 2015, 97(2);189-204
[PubMed:25869574]
[WorldCat.org]
[DOI]
(I p)
Yuh Shiwa, Hirofumi Yoshikawa, Teruo Tanaka, Mitsuo Ogura
Bacillus subtilis degSU operon is regulated by the ClpXP-Spx regulated proteolysis system.
J Biochem: 2015, 157(5);321-30
[PubMed:25433860]
[WorldCat.org]
[DOI]
(I p)
Monica Gupta, K Krishnamurthy Rao
Phosphorylation of DegU is essential for activation of amyE expression in Bacillus subtilis.
J Biosci: 2014, 39(5);747-52
[PubMed:25431404]
[WorldCat.org]
[DOI]
(I p)
Serena Mordini, Cecilia Osera, Simone Marini, Francesco Scavone, Riccardo Bellazzi, Alessandro Galizzi, Cinzia Calvio
The role of SwrA, DegU and P(D3) in fla/che expression in B. subtilis.
PLoS One: 2013, 8(12);e85065
[PubMed:24386445]
[WorldCat.org]
[DOI]
(I e)
Mitsuo Ogura, Hirofumi Yoshikawa, Taku Chibazakura
Regulation of the response regulator gene degU through the binding of SinR/SlrR and exclusion of SinR/SlrR by DegU in Bacillus subtilis.
J Bacteriol: 2014, 196(4);873-81
[PubMed:24317403]
[WorldCat.org]
[DOI]
(I p)
Jia Mun Chan, Sarah B Guttenplan, Daniel B Kearns
Defects in the flagellar motor increase synthesis of poly-γ-glutamate in Bacillus subtilis.
J Bacteriol: 2014, 196(4);740-53
[PubMed:24296669]
[WorldCat.org]
[DOI]
(I p)
Victoria L Marlow, Francesca R Cianfanelli, Michael Porter, Lynne S Cairns, J Kim Dale, Nicola R Stanley-Wall
The prevalence and origin of exoprotease-producing cells in the Bacillus subtilis biofilm.
Microbiology (Reading): 2014, 160(Pt 1);56-66
[PubMed:24149708]
[WorldCat.org]
[DOI]
(I p)
Victoria L Marlow, Michael Porter, Laura Hobley, Taryn B Kiley, Jason R Swedlow, Fordyce A Davidson, Nicola R Stanley-Wall
Phosphorylated DegU manipulates cell fate differentiation in the Bacillus subtilis biofilm.
J Bacteriol: 2014, 196(1);16-27
[PubMed:24123822]
[WorldCat.org]
[DOI]
(I p)
Lynne S Cairns, Victoria L Marlow, Emma Bissett, Adam Ostrowski, Nicola R Stanley-Wall
A mechanical signal transmitted by the flagellum controls signalling in Bacillus subtilis.
Mol Microbiol: 2013, 90(1);6-21
[PubMed:23888912]
[WorldCat.org]
[DOI]
(I p)
Hiroshi Ishii, Teruo Tanaka, Mitsuo Ogura
The Bacillus subtilis response regulator gene degU is positively regulated by CcpA and by catabolite-repressed synthesis of ClpC.
J Bacteriol: 2013, 195(2);193-201
[PubMed:23123903]
[WorldCat.org]
[DOI]
(I p)
Fordyce A Davidson, Chung Seon-Yi, Nicola R Stanley-Wall
Selective heterogeneity in exoprotease production by Bacillus subtilis.
PLoS One: 2012, 7(6);e38574
[PubMed:22745669]
[WorldCat.org]
[DOI]
(I p)
Mitsuo Ogura, Kensuke Tsukahara
SwrA regulates assembly of Bacillus subtilis DegU via its interaction with N-terminal domain of DegU.
J Biochem: 2012, 151(6);643-55
[PubMed:22496484]
[WorldCat.org]
[DOI]
(I p)
Thi-Huyen Do, Yuki Suzuki, Naoki Abe, Jun Kaneko, Yoshifumi Itoh, Keitarou Kimura
Mutations suppressing the loss of DegQ function in Bacillus subtilis (natto) poly-γ-glutamate synthesis.
Appl Environ Microbiol: 2011, 77(23);8249-58
[PubMed:21965392]
[WorldCat.org]
[DOI]
(I p)
Adam Ostrowski, Angela Mehert, Alan Prescott, Taryn B Kiley, Nicola R Stanley-Wall
YuaB functions synergistically with the exopolysaccharide and TasA amyloid fibers to allow biofilm formation by Bacillus subtilis.
J Bacteriol: 2011, 193(18);4821-31
[PubMed:21742882]
[WorldCat.org]
[DOI]
(I p)
Taryn B Kiley, Nicola R Stanley-Wall
Post-translational control of Bacillus subtilis biofilm formation mediated by tyrosine phosphorylation.
Mol Microbiol: 2010, 78(4);947-63
[PubMed:20815827]
[WorldCat.org]
[DOI]
(I p)
Mitsuo Ogura, Kensuke Tsukahara
Autoregulation of the Bacillus subtilis response regulator gene degU is coupled with the proteolysis of DegU-P by ClpCP.
Mol Microbiol: 2010, 75(5);1244-59
[PubMed:20070525]
[WorldCat.org]
[DOI]
(I p)
Taku Ohsawa, Kensuke Tsukahara, Mitsuo Ogura
Bacillus subtilis response regulator DegU is a direct activator of pgsB transcription involved in gamma-poly-glutamic acid synthesis.
Biosci Biotechnol Biochem: 2009, 73(9);2096-102
[PubMed:19734658]
[WorldCat.org]
[DOI]
(I p)
Keitarou Kimura, Lam-Son Phan Tran, Thi-Huyen Do, Yoshifumi Itoh
Expression of the pgsB encoding the poly-gamma-DL-glutamate synthetase of Bacillus subtilis (natto).
Biosci Biotechnol Biochem: 2009, 73(5);1149-55
[PubMed:19420703]
[WorldCat.org]
[DOI]
(I p)
Monica Gupta, Kestur Krishnamurthy Rao
Epr plays a key role in DegU-mediated swarming motility of Bacillus subtilis.
FEMS Microbiol Lett: 2009, 295(2);187-94
[PubMed:19416356]
[WorldCat.org]
[DOI]
(I p)
Cecilia Osera, Giuseppe Amati, Cinzia Calvio, Alessandro Galizzi
SwrAA activates poly-gamma-glutamate synthesis in addition to swarming in Bacillus subtilis.
Microbiology (Reading): 2009, 155(Pt 7);2282-2287
[PubMed:19389763]
[WorldCat.org]
[DOI]
(P p)
Daniel T Verhamme, Ewan J Murray, Nicola R Stanley-Wall
DegU and Spo0A jointly control transcription of two loci required for complex colony development by Bacillus subtilis.
J Bacteriol: 2009, 191(1);100-8
[PubMed:18978066]
[WorldCat.org]
[DOI]
(I p)
Ayako Yasumura, Sadanobu Abe, Teruo Tanaka
Involvement of nitrogen regulation in Bacillus subtilis degU expression.
J Bacteriol: 2008, 190(15);5162-71
[PubMed:18502860]
[WorldCat.org]
[DOI]
(I p)
Jan-Willem Veening, Oleg A Igoshin, Robyn T Eijlander, Reindert Nijland, Leendert W Hamoen, Oscar P Kuipers
Transient heterogeneity in extracellular protease production by Bacillus subtilis.
Mol Syst Biol: 2008, 4;184
[PubMed:18414485]
[WorldCat.org]
[DOI]
(I p)
Kensuke Tsukahara, Mitsuo Ogura
Promoter selectivity of the Bacillus subtilis response regulator DegU, a positive regulator of the fla/che operon and sacB.
BMC Microbiol: 2008, 8;8
[PubMed:18197985]
[WorldCat.org]
[DOI]
(I e)
Kensuke Tsukahara, Mitsuo Ogura
Characterization of DegU-dependent expression of bpr in Bacillus subtilis.
FEMS Microbiol Lett: 2008, 280(1);8-13
[PubMed:18194340]
[WorldCat.org]
[DOI]
(P p)
Kazuo Kobayashi
Gradual activation of the response regulator DegU controls serial expression of genes for flagellum formation and biofilm formation in Bacillus subtilis.
Mol Microbiol: 2007, 66(2);395-409
[PubMed:17850253]
[WorldCat.org]
[DOI]
(P p)
Daniël T Verhamme, Taryn B Kiley, Nicola R Stanley-Wall
DegU co-ordinates multicellular behaviour exhibited by Bacillus subtilis.
Mol Microbiol: 2007, 65(2);554-68
[PubMed:17590234]
[WorldCat.org]
[DOI]
(P p)
Kana Shimane, Mitsuo Ogura
Mutational analysis of the helix-turn-helix region of Bacillus subtilis response regulator DegU, and identification of cis-acting sequences for DegU in the aprE and comK promoters.
J Biochem: 2004, 136(3);387-97
[PubMed:15598897]
[WorldCat.org]
[DOI]
(P p)
Leif Steil, Tamara Hoffmann, Ina Budde, Uwe Völker, Erhard Bremer
Genome-wide transcriptional profiling analysis of adaptation of Bacillus subtilis to high salinity.
J Bacteriol: 2003, 185(21);6358-70
[PubMed:14563871]
[WorldCat.org]
[DOI]
(P p)
Mitsuo Ogura, Kana Shimane, Kei Asai, Naotake Ogasawara, Teruo Tanaka
Binding of response regulator DegU to the aprE promoter is inhibited by RapG, which is counteracted by extracellular PhrG in Bacillus subtilis.
Mol Microbiol: 2003, 49(6);1685-97
[PubMed:12950930]
[WorldCat.org]
[DOI]
(P p)
L W Hamoen, A F Van Werkhoven, G Venema, D Dubnau
The pleiotropic response regulator DegU functions as a priming protein in competence development in Bacillus subtilis.
Proc Natl Acad Sci U S A: 2000, 97(16);9246-51
[PubMed:10908654]
[WorldCat.org]
[DOI]
(P p)
C Fabret, V A Feher, J A Hoch
Two-component signal transduction in Bacillus subtilis: how one organism sees its world.
J Bacteriol: 1999, 181(7);1975-83
[PubMed:10094672]
[WorldCat.org]
[DOI]
(P p)
M Ogura, T Tanaka
Bacillus subtilis DegU acts as a positive regulator for comK expression.
FEBS Lett: 1996, 397(2-3);173-6
[PubMed:8955341]
[WorldCat.org]
[DOI]
(P p)
J Hahn, A Luttinger, D Dubnau
Regulatory inputs for the synthesis of ComK, the competence transcription factor of Bacillus subtilis.
Mol Microbiol: 1996, 21(4);763-75
[PubMed:8878039]
[WorldCat.org]
[DOI]
(P p)
B J Haijema, L W Hamoen, J Kooistra, G Venema, D van Sinderen
Expression of the ATP-dependent deoxyribonuclease of Bacillus subtilis is under competence-mediated control.
Mol Microbiol: 1995, 15(2);203-11
[PubMed:7746142]
[WorldCat.org]
[DOI]
(P p)
K Mukai, M Kawata-Mukai, T Tanaka
Stabilization of phosphorylated Bacillus subtilis DegU by DegR.
J Bacteriol: 1992, 174(24);7954-62
[PubMed:1459944]
[WorldCat.org]
[DOI]
(P p)
M K Dahl, T Msadek, F Kunst, G Rapoport
The phosphorylation state of the DegU response regulator acts as a molecular switch allowing either degradative enzyme synthesis or expression of genetic competence in Bacillus subtilis.
J Biol Chem: 1992, 267(20);14509-14
[PubMed:1321152]
[WorldCat.org]
(P p)
M K Dahl, T Msadek, F Kunst, G Rapoport
Mutational analysis of the Bacillus subtilis DegU regulator and its phosphorylation by the DegS protein kinase.
J Bacteriol: 1991, 173(8);2539-47
[PubMed:1901568]
[WorldCat.org]
[DOI]
(P p)
T Msadek, F Kunst, D Henner, A Klier, G Rapoport, R Dedonder
Signal transduction pathway controlling synthesis of a class of degradative enzymes in Bacillus subtilis: expression of the regulatory genes and analysis of mutations in degS and degU.
J Bacteriol: 1990, 172(2);824-34
[PubMed:1688843]
[WorldCat.org]
[DOI]
(P p)
H Shimotsu, D J Henner
Modulation of Bacillus subtilis levansucrase gene expression by sucrose and regulation of the steady-state mRNA level by sacU and sacQ genes.
J Bacteriol: 1986, 168(1);380-8
[PubMed:2428811]
[WorldCat.org]
[DOI]
(P p)