SipW
Revision as of 14:17, 13 May 2013 by 134.76.70.252 (talk)
- Description: bifunctional signal peptidase I that controls surface-adhered biofilm formation and processes TasA and TapA
| Gene name | sipW |
| Synonyms | yqhE |
| Essential | no |
| Product | signal peptidase I |
| Function | biofilm formation |
| Gene expression levels in SubtiExpress: sipW | |
| Interactions involving this protein in SubtInteract: SipW | |
| Regulation of this protein in SubtiPathways: Biofilm, Protein secretion | |
| MW, pI | 20 kDa, 5.494 |
| Gene length, protein length | 570 bp, 190 aa |
| Immediate neighbours | tasA, tapA |
| Sequences | Protein DNA Advanced_DNA |
Genetic context 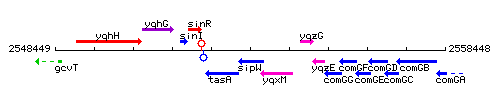
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed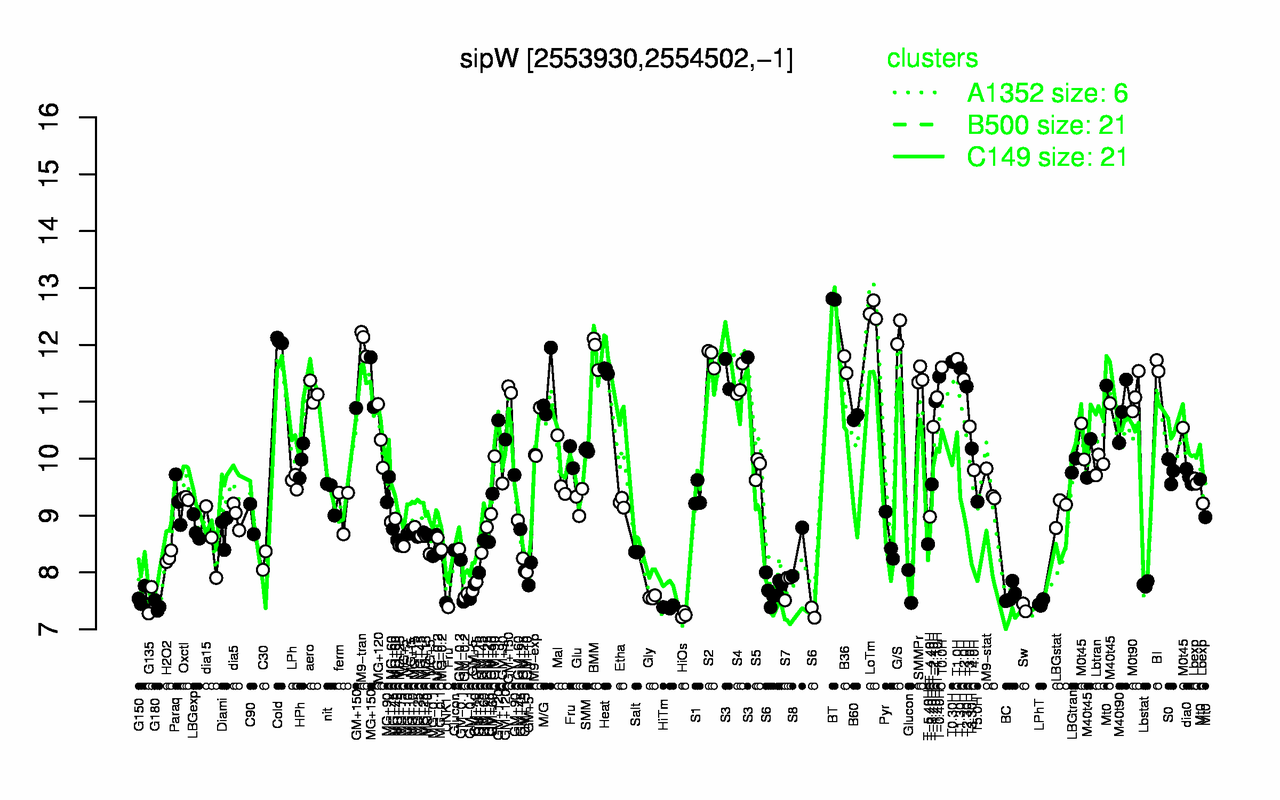
| |
Contents
Categories containing this gene/protein
protein secretion, biofilm formation, membrane proteins
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU24630
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- important for biofilm formation on a solid surface, but not required at an air-liquid interface PubMed
- Cleavage of hydrophobic, N-terminal signal or leader sequences from TasA and TapA PubMed
- Protein family: peptidase S26B family (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Localization:
- membrane PubMed
Database entries
- Structure:
- UniProt: P54506
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Additional information:
Biological materials
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
- Beth Lazazzera, Los Angeles, USA
Your additional remarks
References
Reviews
Additional reviews: PubMed
Adam Driks
Tapping into the biofilm: insights into assembly and disassembly of a novel amyloid fibre in Bacillus subtilis.
Mol Microbiol: 2011, 80(5);1133-6
[PubMed:21488983]
[WorldCat.org]
[DOI]
(I p)
Original publications
Lehnik-Habrink M, Schaffer M, Mäder U, Diethmaier C, Herzberg C, Stülke J RNA processing in Bacillus subtilis: identification of targets of the essential RNase Y. Mol Microbiol. 2011 81(6): 1459-1473. PubMed:21815947
Diethmaier C, Pietack N, Gunka K, Wrede C, Lehnik-Habrink M, Herzberg C, Hübner S, Stülke J A Novel Factor Controlling Bistability in Bacillus subtilis: The YmdB Protein Affects Flagellin Expression and Biofilm Formation. J Bacteriol.: 2011, 193(21):5997-6007. PubMed:21856853