GapA
- Description: Glyceraldehyde 3-phosphate dehydrogenase, NAD-dependent, glycolytic enzyme, forms a transhydrogenation cycle with GapB for balancing of NADPH
| Gene name | gapA |
| Synonyms | |
| Essential | Yes (PubMed) |
| Product | glyceraldehyde 3-phosphate dehydrogenase |
| Function | catabolic enzyme in glycolysis |
| Interactions involving this protein in SubtInteract: GapA | |
| Metabolic function and regulation of this protein in SubtiPathways: Central C-metabolism | |
| MW, pI | 35.7 kDa, 5.03 |
| Gene length, protein length | 1005 bp, 335 amino acids |
| Immediate neighbours | pgk, cggR |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed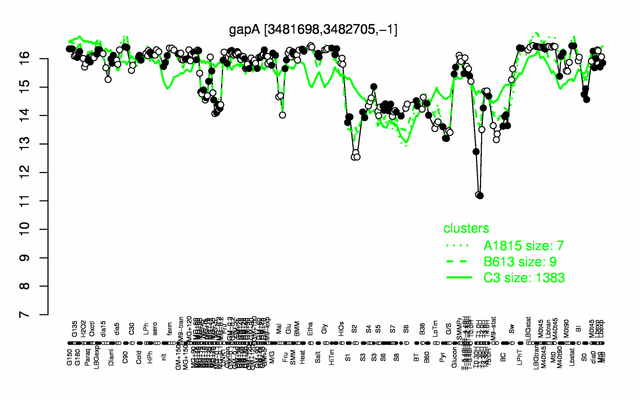
| |
Contents
Categories containing this gene/protein
carbon core metabolism, membrane proteins, phosphoproteins
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU33940
Phenotypes of a mutant
- Essential PubMed
Database entries
- DBTBS entry: [1]
- SubtiList entry:[2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: D-glyceraldehyde 3-phosphate + phosphate + NAD+ = 3-phospho-D-glyceroyl phosphate + NADH (according to Swiss-Prot)
- This reaction is part of the glycolysis.
- Protein family: glyceraldehyde-3-phosphate dehydrogenase family (according to Swiss-Prot)
- Paralogous protein(s): GapB
Extended information on the protein
- Kinetic information: Michaelis-Menten PubMed
- Domains:
- Modification:
- Cofactor(s): NAD+ (do not accept NADP+) PubMed
- Effectors of protein activity:
Database entries
- Structure:
- UniProt: P09124
- KEGG entry: [3]
- E.C. number: 1.2.1.12
Additional information
- GAP dehydrogenases from different sources (incl. Geobacillus stearothermophilus) were shown to cleave RNA (PubMed)
- Moreover, mutations in gapA from B. subtilis can suppress mutations in genes involved in DNA replication (PubMed).
- extensive information on the structure and enzymatic properties of GapA can be found at Proteopedia
Expression and regulation
- Database entries: DBTBS
- Additional information:
- GapA is one of the most abundant proteins in the cell. In the presence of glucose, there are about 25,000 GapA molecules per cell (PubMed).
- The primary mRNAs of the operon are highly unstable. The primary mRNA is subject to processing at the very end of the cggR open reading frame. This results in stable mature gapA and gapA-pgk-tpiA-pgm-eno mRNAs. PubMed The processing event requires the RNase Y PubMed.
- The accumulation of the cggR-gapA mRNA is strongly dependent on the presence of the YkzW peptide, due to stabilization of the mRNA PubMed.
- the mRNA is substantially stabilized upon depletion of RNase Y PubMed
Biological materials
- Mutant:
- GP592 (gapA::cat), available in Stülke lab
- GP597 (gapA::erm), available in Stülke lab
- GP703 (gapA::cat gapB::spec), available in Stülke lab
- GM1501 (under p(spac) control), available in Stephane Aymerich lab
- Expression vector:
- GFP fusion:
- two-hybrid system: B. pertussis adenylate cyclase-based bacterial two hybrid system (BACTH), available in Stülke lab
- Antibody: available in Stülke lab
Labs working on this gene/protein
Stephane Aymerich, Microbiology and Molecular Genetics, INRA Paris-Grignon, France
Jörg Stülke, University of Göttingen, Germany homepage
Your additional remarks
References
Lehnik-Habrink M, Schaffer M, Mäder U, Diethmaier C, Herzberg C, Stülke J RNA processing in Bacillus subtilis: identification of targets of the essential RNase Y. Mol Microbiol. 2011 81(6): 1459-1473. PubMed:21815947
Bui Khanh Chi, Katrin Gronau, Ulrike Mäder, Bernd Hessling, Dörte Becher, Haike Antelmann
S-bacillithiolation protects against hypochlorite stress in Bacillus subtilis as revealed by transcriptomics and redox proteomics.
Mol Cell Proteomics: 2011, 10(11);M111.009506
[PubMed:21749987]
[WorldCat.org]
[DOI]
(I p)
Matthias Gimpel, Nadja Heidrich, Ulrike Mäder, Hans Krügel, Sabine Brantl
A dual-function sRNA from B. subtilis: SR1 acts as a peptide encoding mRNA on the gapA operon.
Mol Microbiol: 2010, 76(4);990-1009
[PubMed:20444087]
[WorldCat.org]
[DOI]
(I p)
Fabian M Commichau, Fabian M Rothe, Christina Herzberg, Eva Wagner, Daniel Hellwig, Martin Lehnik-Habrink, Elke Hammer, Uwe Völker, Jörg Stülke
Novel activities of glycolytic enzymes in Bacillus subtilis: interactions with essential proteins involved in mRNA processing.
Mol Cell Proteomics: 2009, 8(6);1350-60
[PubMed:19193632]
[WorldCat.org]
[DOI]
(I p)
Manuel Liebeke, Dierk-Christoph Pöther, Nguyen van Duy, Dirk Albrecht, Dörte Becher, Falko Hochgräfe, Michael Lalk, Michael Hecker, Haike Antelmann
Depletion of thiol-containing proteins in response to quinones in Bacillus subtilis.
Mol Microbiol: 2008, 69(6);1513-29
[PubMed:18673455]
[WorldCat.org]
[DOI]
(I p)
Christine Eymann, Dörte Becher, Jörg Bernhardt, Katrin Gronau, Anja Klutzny, Michael Hecker
Dynamics of protein phosphorylation on Ser/Thr/Tyr in Bacillus subtilis.
Proteomics: 2007, 7(19);3509-26
[PubMed:17726680]
[WorldCat.org]
[DOI]
(P p)
Laurent Jannière, Danielle Canceill, Catherine Suski, Sophie Kanga, Bérengère Dalmais, Roxane Lestini, Anne-Françoise Monnier, Jérôme Chapuis, Alexander Bolotin, Marina Titok, Emmanuelle Le Chatelier, S Dusko Ehrlich
Genetic evidence for a link between glycolysis and DNA replication.
PLoS One: 2007, 2(5);e447
[PubMed:17505547]
[WorldCat.org]
[DOI]
(I e)
Boris Macek, Ivan Mijakovic, Jesper V Olsen, Florian Gnad, Chanchal Kumar, Peter R Jensen, Matthias Mann
The serine/threonine/tyrosine phosphoproteome of the model bacterium Bacillus subtilis.
Mol Cell Proteomics: 2007, 6(4);697-707
[PubMed:17218307]
[WorldCat.org]
[DOI]
(P p)
Frédérique Pompeo, Jennifer Luciano, Anne Galinier
Interaction of GapA with HPr and its homologue, Crh: Novel levels of regulation of a key step of glycolysis in Bacillus subtilis?
J Bacteriol: 2007, 189(3);1154-7
[PubMed:17142398]
[WorldCat.org]
[DOI]
(P p)
Helena B Thomaides, Ella J Davison, Lisa Burston, Hazel Johnson, David R Brown, Alison C Hunt, Jeffery Errington, Lloyd Czaplewski
Essential bacterial functions encoded by gene pairs.
J Bacteriol: 2007, 189(2);591-602
[PubMed:17114254]
[WorldCat.org]
[DOI]
(P p)
Jean-Christophe Meile, Ling Juan Wu, S Dusko Ehrlich, Jeff Errington, Philippe Noirot
Systematic localisation of proteins fused to the green fluorescent protein in Bacillus subtilis: identification of new proteins at the DNA replication factory.
Proteomics: 2006, 6(7);2135-46
[PubMed:16479537]
[WorldCat.org]
[DOI]
(P p)
Hans-Matti Blencke, Georg Homuth, Holger Ludwig, Ulrike Mäder, Michael Hecker, Jörg Stülke
Transcriptional profiling of gene expression in response to glucose in Bacillus subtilis: regulation of the central metabolic pathways.
Metab Eng: 2003, 5(2);133-49
[PubMed:12850135]
[WorldCat.org]
[DOI]
(P p)
Christoph Meinken, Hans-Matti Blencke, Holger Ludwig, Jörg Stülke
Expression of the glycolytic gapA operon in Bacillus subtilis: differential syntheses of proteins encoded by the operon.
Microbiology (Reading): 2003, 149(Pt 3);751-761
[PubMed:12634343]
[WorldCat.org]
[DOI]
(P p)
Thierry Doan, Stéphane Aymerich
Regulation of the central glycolytic genes in Bacillus subtilis: binding of the repressor CggR to its single DNA target sequence is modulated by fructose-1,6-bisphosphate.
Mol Microbiol: 2003, 47(6);1709-21
[PubMed:12622823]
[WorldCat.org]
[DOI]
(P p)
Elena Evguenieva-Hackenberg, Emile Schiltz, Gabriele Klug
Dehydrogenases from all three domains of life cleave RNA.
J Biol Chem: 2002, 277(48);46145-50
[PubMed:12359717]
[WorldCat.org]
[DOI]
(P p)
Holger Ludwig, Nicole Rebhan, Hans-Matti Blencke, Matthias Merzbacher, Jörg Stülke
Control of the glycolytic gapA operon by the catabolite control protein A in Bacillus subtilis: a novel mechanism of CcpA-mediated regulation.
Mol Microbiol: 2002, 45(2);543-53
[PubMed:12123463]
[WorldCat.org]
[DOI]
(P p)
H Ludwig, G Homuth, M Schmalisch, F M Dyka, M Hecker, J Stülke
Transcription of glycolytic genes and operons in Bacillus subtilis: evidence for the presence of multiple levels of control of the gapA operon.
Mol Microbiol: 2001, 41(2);409-22
[PubMed:11489127]
[WorldCat.org]
[DOI]
(P p)
S Fillinger, S Boschi-Muller, S Azza, E Dervyn, G Branlant, S Aymerich
Two glyceraldehyde-3-phosphate dehydrogenases with opposite physiological roles in a nonphotosynthetic bacterium.
J Biol Chem: 2000, 275(19);14031-7
[PubMed:10799476]
[WorldCat.org]
[DOI]
(P p)
S Tobisch, D Zühlke, J Bernhardt, J Stülke, M Hecker
Role of CcpA in regulation of the central pathways of carbon catabolism in Bacillus subtilis.
J Bacteriol: 1999, 181(22);6996-7004
[PubMed:10559165]
[WorldCat.org]
[DOI]
(P p)