| Gene name
|
eno
|
| Synonyms |
|
| Essential |
no
|
| Product |
enolase
|
| Function |
enzyme in glycolysis/ gluconeogenesis
|
| Interactions involving this protein in SubtInteract: Eno
|
Metabolic function and regulation of this protein in SubtiPathways:
Central C-metabolism
|
| MW, pI |
46,4 kDa, 4.49
|
| Gene length, protein length |
1290 bp, 430 amino acids
|
| Immediate neighbours |
yvbK, pgm
|
Get the DNA and protein sequences
(Barbe et al., 2009)
|
Genetic context
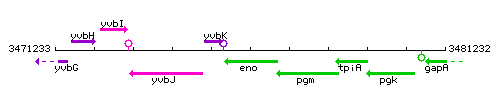
|
|
|
Categories containing this gene/protein
carbon core metabolism,
membrane proteins,
phosphoproteins,
universally conserved proteins
This gene is a member of the following regulons
CggR regulon
The gene
Basic information
Phenotypes of a mutant
- no growth on LB, requires glucose and malate
- essential according to Kobayashi et al. on LB PubMed
Database entries
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: 2-phospho-D-glycerate = phosphoenolpyruvate + H2O (according to Swiss-Prot) 2-phospho-D-glycerate = phosphoenolpyruvate + H(2)O
- Protein family: enolase family (according to Swiss-Prot)
Extended information on the protein
- Kinetic information: reversible Michaelis-Menten PubMed
- Domains:
- substrate binding domain (366–369)
- Modification: phosphorylation on Thr-141 AND Ser-259 AND Tyr-281 AND Ser-325 PubMed, PubMed, PubMed
- Effectors of protein activity:
Database entries
Additional information
Expression and regulation
- Regulatory mechanism: transcription repression by CggR PubMed
Biological materials
- Mutant:
- GP594 (eno::cat), available in Stülke lab
- GP599 (eno::erm), available in Stülke lab
- GP698 (eno-pgm::cat), available in Stülke lab
- Expression vector:
- pGP1426 (expression of eno in B. subtilis, in pBQ200), available in Stülke lab
- pGP1500 (expression of pgm and eno in B. subtilis, in pBQ200), available in Stülke lab
- pGP563 (N-terminal His-tag, in pWH844), available in Stülke lab
- pGP1276 (N-terminal Strep-tag, purification from E. coli, in pGP172), available in Stülke lab
- pGP93 (N-terminal Strep-tag, purification from B. subtilis, for SPINE, in pGP380), available in Stülke lab
- GP1215 (eno-Strep (spc)), purification from B. subtilis, for SPINE, available in Stülke lab
- GFP fusion: pHT315-yfp-eno, available in Mijakovic lab
- two-hybrid system: B. pertussis adenylate cyclase-based bacterial two hybrid system (BACTH), available in Stülke lab
- FLAG-tag construct: GP1214 (spc, based on pGP1331), available in the Stülke lab
- Antibody: available in Stülke lab
Labs working on this gene/protein
Jörg Stülke, University of Göttingen, Germany
Homepage
References
Reviews
Subcellular localization of enolase
Additional publications: PubMed
Other original publications
