SigM
- Description: RNA polymerase ECF-type sigma factor SigM
| Gene name | sigM |
| Synonyms | yhdM |
| Essential | no |
| Product | RNA polymerase ECF-type sigma factor SigM |
| Function | responsible for intrinsic resistance against beta-lactam antibiotics |
| Gene expression levels in SubtiExpress: sigM | |
| Interactions involving this protein in SubtInteract: SigM | |
| MW, pI | 19 kDa, 6.77 |
| Gene length, protein length | 489 bp, 163 aa |
| Immediate neighbours | yhdL, yhdN |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 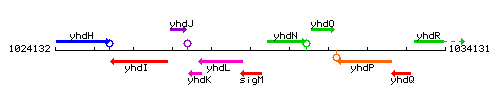
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed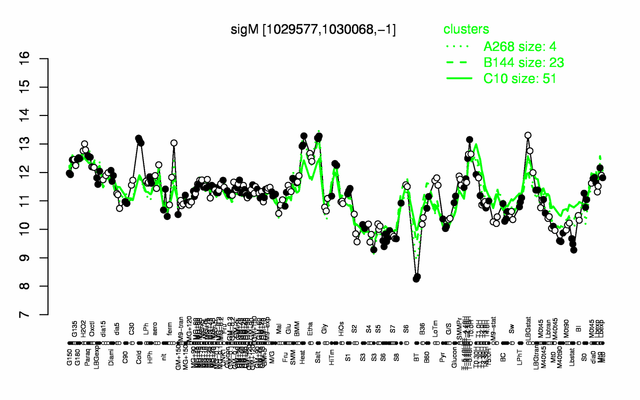
| |
Contents
Categories containing this gene/protein
transcription, sigma factors and their control, cell envelope stress proteins (controlled by SigM, V, W, X, Y), resistance against toxins/ antibiotics
This gene is a member of the following regulons
The SigM regulon
The gene
Basic information
- Locus tag: BSU09520
Phenotypes of a mutant
- SigM is essential for growth and survival in nutrient broth (NB) containing 1.4 M NaCl PubMed
- sigM mutants form aberrantly shaped cells, which swell and lyse spontaneously during growth in NB medium containing increased levels (0.35-0.7 M) of a wide range of different salts PubMed
- increased sensitivity towards beta-lactam antibiotics PubMed
Database entries
- BsubCyc: BSU09520
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family: sigma-70 factor family, ECF subfamily (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
Database entries
- BsubCyc: BSU09520
- Structure:
- UniProt: O07582
- KEGG entry: [3]
- E.C. number:
Additional information
- Expression of the SigM regulon in increased in ugtP mutants PubMed
Expression and regulation
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant:
- 1A906 (sigM::kan), PubMed, available at BGSC
- BP129 (sigM::aphA3), available in Fabian Commichau's lab
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
John Helmann, Cornell University, USA Homepage
Your additional remarks
References
Reviews
The SigM regulon
Other original publications
Hiromi Inoue, Daisuke Suzuki, Kei Asai
A putative bactoprenol glycosyltransferase, CsbB, in Bacillus subtilis activates SigM in the absence of co-transcribed YfhO.
Biochem Biophys Res Commun: 2013, 436(1);6-11
[PubMed:23632331]
[WorldCat.org]
[DOI]
(I p)
Michihiro Hashimoto, Takahiro Seki, Satoshi Matsuoka, Hiroshi Hara, Kei Asai, Yoshito Sadaie, Kouji Matsumoto
Induction of extracytoplasmic function sigma factors in Bacillus subtilis cells with defects in lipoteichoic acid synthesis.
Microbiology (Reading): 2013, 159(Pt 1);23-35
[PubMed:23103977]
[WorldCat.org]
[DOI]
(I p)
Yong Heon Lee, Ki Hyun Nam, John D Helmann
A mutation of the RNA polymerase β' subunit (rpoC) confers cephalosporin resistance in Bacillus subtilis.
Antimicrob Agents Chemother: 2013, 57(1);56-65
[PubMed:23070162]
[WorldCat.org]
[DOI]
(I p)
Satoshi Matsuoka, Minako Chiba, Yu Tanimura, Michihiro Hashimoto, Hiroshi Hara, Kouji Matsumoto
Abnormal morphology of Bacillus subtilis ugtP mutant cells lacking glucolipids.
Genes Genet Syst: 2011, 86(5);295-304
[PubMed:22362028]
[WorldCat.org]
[DOI]
(I p)
Yun Luo, John D Helmann
Analysis of the role of Bacillus subtilis σ(M) in β-lactam resistance reveals an essential role for c-di-AMP in peptidoglycan homeostasis.
Mol Microbiol: 2012, 83(3);623-39
[PubMed:22211522]
[WorldCat.org]
[DOI]
(I p)
Yun Luo, Kei Asai, Yoshito Sadaie, John D Helmann
Transcriptomic and phenotypic characterization of a Bacillus subtilis strain without extracytoplasmic function σ factors.
J Bacteriol: 2010, 192(21);5736-45
[PubMed:20817771]
[WorldCat.org]
[DOI]
(I p)
Andriansjah Rukmana, Takuya Morimoto, Hiroki Takahashi, Giyanto, Naotake Ogasawara
Assessment of transcriptional responses of Bacillus subtilis cells to the antibiotic enduracidin, which interferes with cell wall synthesis, using a high-density tiling chip.
Genes Genet Syst: 2009, 84(4);253-67
[PubMed:20057163]
[WorldCat.org]
[DOI]
(P p)
Michihiro Hashimoto, Hiroaki Takahashi, Yoshinori Hara, Hiroshi Hara, Kei Asai, Yoshito Sadaie, Kouji Matsumoto
Induction of extracytoplasmic function sigma factors in Bacillus subtilis cells with membranes of reduced phosphatidylglycerol content.
Genes Genet Syst: 2009, 84(3);191-8
[PubMed:19745567]
[WorldCat.org]
[DOI]
(P p)
Yun Luo, John D Helmann
Extracytoplasmic function sigma factors with overlapping promoter specificity regulate sublancin production in Bacillus subtilis.
J Bacteriol: 2009, 191(15);4951-8
[PubMed:19465659]
[WorldCat.org]
[DOI]
(I p)
Warawan Eiamphungporn, John D Helmann
Extracytoplasmic function sigma factors regulate expression of the Bacillus subtilis yabE gene via a cis-acting antisense RNA.
J Bacteriol: 2009, 191(3);1101-5
[PubMed:19047346]
[WorldCat.org]
[DOI]
(I p)
Letal I Salzberg, John D Helmann
Phenotypic and transcriptomic characterization of Bacillus subtilis mutants with grossly altered membrane composition.
J Bacteriol: 2008, 190(23);7797-807
[PubMed:18820022]
[WorldCat.org]
[DOI]
(I p)
Eva Rietkötter, Diana Hoyer, Thorsten Mascher
Bacitracin sensing in Bacillus subtilis.
Mol Microbiol: 2008, 68(3);768-85
[PubMed:18394148]
[WorldCat.org]
[DOI]
(I p)
Kathrin Minnig, Vladimir Lazarevic, Blazenka Soldo, Catherine Mauël
Analysis of teichoic acid biosynthesis regulation reveals that the extracytoplasmic function sigma factor sigmaM is induced by phosphate depletion in Bacillus subtilis W23.
Microbiology (Reading): 2005, 151(Pt 9);3041-3049
[PubMed:16151214]
[WorldCat.org]
[DOI]
(P p)
Min Cao, Charles M Moore, John D Helmann
Bacillus subtilis paraquat resistance is directed by sigmaM, an extracytoplasmic function sigma factor, and is conferred by YqjL and BcrC.
J Bacteriol: 2005, 187(9);2948-56
[PubMed:15838020]
[WorldCat.org]
[DOI]
(P p)
Mika Yoshimura, Kei Asai, Yoshito Sadaie, Hirofumi Yoshikawa
Interaction of Bacillus subtilis extracytoplasmic function (ECF) sigma factors with the N-terminal regions of their potential anti-sigma factors.
Microbiology (Reading): 2004, 150(Pt 3);591-599
[PubMed:14993308]
[WorldCat.org]
[DOI]
(P p)
Thorsten Mascher, Neil G Margulis, Tao Wang, Rick W Ye, John D Helmann
Cell wall stress responses in Bacillus subtilis: the regulatory network of the bacitracin stimulon.
Mol Microbiol: 2003, 50(5);1591-604
[PubMed:14651641]
[WorldCat.org]
[DOI]
(P p)
Penny D Thackray, Anne Moir
SigM, an extracytoplasmic function sigma factor of Bacillus subtilis, is activated in response to cell wall antibiotics, ethanol, heat, acid, and superoxide stress.
J Bacteriol: 2003, 185(12);3491-8
[PubMed:12775685]
[WorldCat.org]
[DOI]
(P p)
Min Cao, John D Helmann
Regulation of the Bacillus subtilis bcrC bacitracin resistance gene by two extracytoplasmic function sigma factors.
J Bacteriol: 2002, 184(22);6123-9
[PubMed:12399481]
[WorldCat.org]
[DOI]
(P p)
Min Cao, Tao Wang, Rick Ye, John D Helmann
Antibiotics that inhibit cell wall biosynthesis induce expression of the Bacillus subtilis sigma(W) and sigma(M) regulons.
Mol Microbiol: 2002, 45(5);1267-76
[PubMed:12207695]
[WorldCat.org]
[DOI]
(P p)
M J Horsburgh, A Moir
Sigma M, an ECF RNA polymerase sigma factor of Bacillus subtilis 168, is essential for growth and survival in high concentrations of salt.
Mol Microbiol: 1999, 32(1);41-50
[PubMed:10216858]
[WorldCat.org]
[DOI]
(P p)