Difference between revisions of "Sandbox"
| Line 1: | Line 1: | ||
| − | * '''Description:''' subunit of | + | * '''Description:''' small subunit of glutamate synthase<br/><br/> |
{| align="right" border="1" cellpadding="2" | {| align="right" border="1" cellpadding="2" | ||
|- | |- | ||
|style="background:#ABCDEF;" align="center"|'''Gene name''' | |style="background:#ABCDEF;" align="center"|'''Gene name''' | ||
| − | |'' | + | |''gltB'' |
|- | |- | ||
| − | |style="background:#ABCDEF;" align="center"| '''Synonyms''' || '' | + | |style="background:#ABCDEF;" align="center"| '''Synonyms''' || '' '' |
|- | |- | ||
| − | |style="background:#ABCDEF;" align="center"| '''Essential''' || | + | |style="background:#ABCDEF;" align="center"| '''Essential''' || no |
|- | |- | ||
| − | |style="background:#ABCDEF;" align="center"| '''Product''' || subunit | + | |style="background:#ABCDEF;" align="center"| '''Product''' || glutamate synthase (small subunit) |
|- | |- | ||
| − | |style="background:#ABCDEF;" align="center"|'''Function''' || | + | |style="background:#ABCDEF;" align="center"|'''Function''' || glutamate biosynthesis |
|- | |- | ||
| − | |style="background:#ABCDEF;" align="center"| '''MW, pI''' || | + | |style="background:#ABCDEF;" align="center"| '''MW, pI''' || 54.6 kDa, 7.69 |
|- | |- | ||
| − | |style="background:#ABCDEF;" align="center"| '''Gene length, protein length''' || | + | |style="background:#ABCDEF;" align="center"| '''Gene length, protein length''' || 1479 bp, 493 amino acids |
|- | |- | ||
| − | |style="background:#ABCDEF;" align="center"|'''Immediate neighbours''' || ''[[ | + | |style="background:#ABCDEF;" align="center"|'''Immediate neighbours''' || ''[[gltA]]'', ''[[yogA]]'' |
|- | |- | ||
| − | |colspan="2" style="background:#FAF8CC;" align="center"|'''Get the DNA and protein [http://srs.ebi.ac.uk/srsbin/cgi-bin/wgetz?-e+[EMBLCDS: | + | |colspan="2" style="background:#FAF8CC;" align="center"|'''Get the DNA and protein [http://srs.ebi.ac.uk/srsbin/cgi-bin/wgetz?-e+[EMBLCDS:CAB13727]+-newId sequences] <br/> (Barbe ''et al.'', 2009)''' |
|- | |- | ||
|- | |- | ||
|- | |- | ||
| − | |colspan="2" | '''Genetic context''' <br/> [[Image: | + | |colspan="2" | '''Genetic context''' <br/> [[Image:gltB_context.gif]] |
<div align="right"> <small>This image was kindly provided by [http://genolist.pasteur.fr/SubtiList/ SubtiList]</small></div> | <div align="right"> <small>This image was kindly provided by [http://genolist.pasteur.fr/SubtiList/ SubtiList]</small></div> | ||
|- | |- | ||
| Line 31: | Line 31: | ||
__TOC__ | __TOC__ | ||
| − | <br/><br/> | + | <br/><br/> |
| + | |||
=The gene= | =The gene= | ||
| Line 37: | Line 38: | ||
=== Basic information === | === Basic information === | ||
| − | * '''Coordinates:''' | + | * '''Coordinates:''' 2007785 - 2009263 |
===Phenotypes of a mutant === | ===Phenotypes of a mutant === | ||
| − | + | auxotrophic for glutamate | |
=== Database entries === | === Database entries === | ||
| − | * '''DBTBS entry:''' [http://dbtbs.hgc.jp/COG/prom/ | + | * '''DBTBS entry:''' [http://dbtbs.hgc.jp/COG/prom/gltAB.html] |
| − | * '''SubtiList entry:''' [http://genolist.pasteur.fr/SubtiList/genome.cgi?gene_detail+ | + | * '''SubtiList entry:'''[http://genolist.pasteur.fr/SubtiList/genome.cgi?gene_detail+BG12594] |
=== Additional information=== | === Additional information=== | ||
| − | |||
=The protein= | =The protein= | ||
| Line 56: | Line 56: | ||
=== Basic information/ Evolution === | === Basic information/ Evolution === | ||
| − | * '''Catalyzed reaction/ biological activity:''' | + | * '''Catalyzed reaction/ biological activity:''' 2 L-glutamate + NADP(+) <=> L-glutamine + 2-oxoglutarate + NADPH |
| − | * '''Protein family:''' | + | * '''Protein family:''' glutamate synthase family. |
| − | * '''Paralogous protein(s):''' | + | * '''Paralogous protein(s):''' none |
=== Extended information on the protein === | === Extended information on the protein === | ||
| Line 67: | Line 67: | ||
* '''Domains:''' | * '''Domains:''' | ||
| + | ** nucleotide binding domain (NADP) (299–313) | ||
* '''Modification:''' | * '''Modification:''' | ||
| Line 76: | Line 77: | ||
* '''Interactions:''' | * '''Interactions:''' | ||
| − | * '''Localization:''' Cytoplasm | + | * '''Localization:''' Cytoplasm |
=== Database entries === | === Database entries === | ||
| Line 82: | Line 83: | ||
* '''Structure:''' | * '''Structure:''' | ||
| − | * '''Swiss prot entry:''' | + | * '''Swiss prot entry:''' [http://www.uniprot.org/uniprot/O34399] |
| − | * '''KEGG entry:''' [http://www.genome.jp/dbget-bin/www_bget?bsu+ | + | * '''KEGG entry:''' [http://www.genome.jp/dbget-bin/www_bget?bsu+BSU18440] |
| − | * '''E.C. number:''' | + | * '''E.C. number:''' [http://www.expasy.ch/cgi-bin/get-enzyme-entry?1.4.1.13 1.4.1.13] |
=== Additional information=== | === Additional information=== | ||
| + | |||
=Expression and regulation= | =Expression and regulation= | ||
| − | * '''Operon:''' | + | * '''Operon:''' ''[[gltA]]-[[gltB]]'' |
| − | * '''[[Sigma factor]]:''' | + | * '''[[Sigma factor]]:''' [[SigA]] |
| − | * '''Regulation:''' | + | * '''Regulation:''' see ''[[gltA]]'' |
| − | + | * '''Additional information:''' | |
| − | |||
| − | * '''Additional information:''' | ||
=Biological materials = | =Biological materials = | ||
| − | * '''Mutant:''' | + | * '''Mutant:''' GP807 (del ''gltAB''::''tet''), GP736 (spc), available in [[Stülke]] lab |
| − | * '''Expression vector:''' | + | * '''Expression vector:''' pGP1119 (in [[pGP380]], for SPINE, expression in ''B. subtilis''), available in [[Stülke]] lab |
| − | + | ||
| − | * '''lacZ fusion:''' | + | * '''lacZ fusion:''' see ''[[gltA]]'' |
* '''GFP fusion:''' | * '''GFP fusion:''' | ||
| − | |||
| − | |||
* '''Antibody:''' | * '''Antibody:''' | ||
=Labs working on this gene/protein= | =Labs working on this gene/protein= | ||
| + | |||
| + | [[Linc Sonenshein|Linc Sonenshein]], Tufts University, Boston, MA, USA [http://www.tufts.edu/sackler/microbiology/faculty/sonenshein/index.html Homepage] | ||
| + | |||
| + | [[Stülke|Jörg Stülke]], University of Göttingen, Germany | ||
| + | [http://wwwuser.gwdg.de/~genmibio/stuelke.html Homepage] | ||
=Your additional remarks= | =Your additional remarks= | ||
| Line 122: | Line 125: | ||
=References= | =References= | ||
| − | # | + | # Yoshida K, et al. (2003) Identification of additional TnrA-regulated genes of Bacillus subtilis associated with a TnrA box. ''Mol Microbiol'' '''49(1):''' 157-65. [http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=+12823818 PubMed] |
| − | # | + | # Belitsky, B. R., and Sonenshein, A. L. (1995) Mutations in GltC that increase ''Bacillus subtilis gltA'' expression. J Bacteriol 177: 5696-5700.[http://www.ncbi.nlm.nih.gov/sites/entrez/7559360 PubMed] |
| + | # Belitsky, B. R., and Sonenshein, A. L. (2004) Modulation of activity of ''Bacillus subtilis'' regulatory proteins GltC and TnrA by glutamate dehydrogenase. J Bacteriol 186: 3399-3407.[http://www.ncbi.nlm.nih.gov/sites/entrez/15150225 PubMed] | ||
| + | # Bohannon, D. E., and Sonenshein, A. L. (1989) Positive regulation of glutamate biosynthesis in ''Bacillus subtilis''. J Bacteriol 171: 4718-4727.[http://www.ncbi.nlm.nih.gov/sites/entrez/2548995 PubMed] | ||
| + | # Commichau, F. M., Wacker, I., Schleider, J., Blencke, H.-M., Reif, I., Tripal, P., and Stülke, J. (2007) Characterization of ''Bacillus subtilis'' mutants with carbon source-independent glutamate biosynthesis. J Mol Microbiol Biotechnol 12: 106-113. [http://www.ncbi.nlm.nih.gov/sites/entrez/17183217 PubMed] | ||
| + | # Commichau, F. M., Herzberg, C., Tripal, P., Valerius, O., and Stülke, J. (2007) A regulatory protein-protein interaction governs glutamate biosynthesis in ''Bacillus subtilis'': The glutamate dehydrogenase RocG moonlights in controlling the transcription factor GltC. Mol Microbiol 65: 642-654. [http://www.ncbi.nlm.nih.gov/sites/entrez/17608797 PubMed] | ||
| + | # Picossi, S., Belitsky, B. R., and Sonenshein, A. L. (2007) Molecular mechanism of the regulation of ''Bacillus subtilis gltAB'' expression by GltC. J Mol Biol 365: 1298-1313. [http://www.ncbi.nlm.nih.gov/sites/entrez/17134717 PubMed] | ||
| + | # Wacker, I., Ludwig, H., Reif, I., Blencke, H. M., Detsch, C., and Stülke, J. (2003) The regulatory link between carbon and nitrogen metabolism in ''Bacillus subtilis'': regulation of the ''gltAB'' operon by the catabolite control protein CcpA. Microbiology 149: 3001-3009.[http://www.ncbi.nlm.nih.gov/sites/entrez/14523131 PubMed] | ||
| + | # Belitsky, B. R., Wray, L. V., Jr., Fisher, S. H., Bohannon, D. E. & Sonenshein, A. L. (2000). Role of TnrA in nitrogen source-dependent repression of ''Bacillus subtilis'' glutamate synthase gene expression. J Bacteriol 182, 5939-5947. [http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=+11029411 PubMed] | ||
| + | # Commichau, F. M., Gunka, K., Landmann, J. J. & Stülke, J. (2008) Glutamate metabolism in ''Bacillus subtilis'': Gene expression and enzyme activities evolved to avoid futile cycles and to allow rapid responses to perturbations in the system. J. Bacteriol. 190: 3557-3564. [http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=+18326565 PubMed] | ||
Revision as of 02:27, 30 April 2009
- Description: small subunit of glutamate synthase
| Gene name | gltB |
| Synonyms | |
| Essential | no |
| Product | glutamate synthase (small subunit) |
| Function | glutamate biosynthesis |
| MW, pI | 54.6 kDa, 7.69 |
| Gene length, protein length | 1479 bp, 493 amino acids |
| Immediate neighbours | gltA, yogA |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 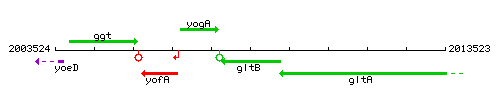
This image was kindly provided by SubtiList
| |
Contents
The gene
Basic information
- Coordinates: 2007785 - 2009263
Phenotypes of a mutant
auxotrophic for glutamate
Database entries
- DBTBS entry: [1]
- SubtiList entry:[2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: 2 L-glutamate + NADP(+) <=> L-glutamine + 2-oxoglutarate + NADPH
- Protein family: glutamate synthase family.
- Paralogous protein(s): none
Extended information on the protein
- Kinetic information:
- Domains:
- nucleotide binding domain (NADP) (299–313)
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Interactions:
- Localization: Cytoplasm
Database entries
- Structure:
- Swiss prot entry: [3]
- KEGG entry: [4]
- E.C. number: 1.4.1.13
Additional information
Expression and regulation
- Regulation: see gltA
- Additional information:
Biological materials
- Mutant: GP807 (del gltAB::tet), GP736 (spc), available in Stülke lab
- Expression vector: pGP1119 (in pGP380, for SPINE, expression in B. subtilis), available in Stülke lab
- lacZ fusion: see gltA
- GFP fusion:
- Antibody:
Labs working on this gene/protein
Linc Sonenshein, Tufts University, Boston, MA, USA Homepage
Jörg Stülke, University of Göttingen, Germany Homepage
Your additional remarks
References
- Yoshida K, et al. (2003) Identification of additional TnrA-regulated genes of Bacillus subtilis associated with a TnrA box. Mol Microbiol 49(1): 157-65. PubMed
- Belitsky, B. R., and Sonenshein, A. L. (1995) Mutations in GltC that increase Bacillus subtilis gltA expression. J Bacteriol 177: 5696-5700.PubMed
- Belitsky, B. R., and Sonenshein, A. L. (2004) Modulation of activity of Bacillus subtilis regulatory proteins GltC and TnrA by glutamate dehydrogenase. J Bacteriol 186: 3399-3407.PubMed
- Bohannon, D. E., and Sonenshein, A. L. (1989) Positive regulation of glutamate biosynthesis in Bacillus subtilis. J Bacteriol 171: 4718-4727.PubMed
- Commichau, F. M., Wacker, I., Schleider, J., Blencke, H.-M., Reif, I., Tripal, P., and Stülke, J. (2007) Characterization of Bacillus subtilis mutants with carbon source-independent glutamate biosynthesis. J Mol Microbiol Biotechnol 12: 106-113. PubMed
- Commichau, F. M., Herzberg, C., Tripal, P., Valerius, O., and Stülke, J. (2007) A regulatory protein-protein interaction governs glutamate biosynthesis in Bacillus subtilis: The glutamate dehydrogenase RocG moonlights in controlling the transcription factor GltC. Mol Microbiol 65: 642-654. PubMed
- Picossi, S., Belitsky, B. R., and Sonenshein, A. L. (2007) Molecular mechanism of the regulation of Bacillus subtilis gltAB expression by GltC. J Mol Biol 365: 1298-1313. PubMed
- Wacker, I., Ludwig, H., Reif, I., Blencke, H. M., Detsch, C., and Stülke, J. (2003) The regulatory link between carbon and nitrogen metabolism in Bacillus subtilis: regulation of the gltAB operon by the catabolite control protein CcpA. Microbiology 149: 3001-3009.PubMed
- Belitsky, B. R., Wray, L. V., Jr., Fisher, S. H., Bohannon, D. E. & Sonenshein, A. L. (2000). Role of TnrA in nitrogen source-dependent repression of Bacillus subtilis glutamate synthase gene expression. J Bacteriol 182, 5939-5947. PubMed
- Commichau, F. M., Gunka, K., Landmann, J. J. & Stülke, J. (2008) Glutamate metabolism in Bacillus subtilis: Gene expression and enzyme activities evolved to avoid futile cycles and to allow rapid responses to perturbations in the system. J. Bacteriol. 190: 3557-3564. PubMed