Difference between revisions of "TrpE"
| Line 39: | Line 39: | ||
<br/><br/><br/><br/> | <br/><br/><br/><br/> | ||
<br/><br/><br/><br/> | <br/><br/><br/><br/> | ||
| − | + | <br/><br/> | |
| − | |||
| − | |||
| − | |||
| − | |||
= [[Categories]] containing this gene/protein = | = [[Categories]] containing this gene/protein = | ||
| Line 66: | Line 62: | ||
=== Additional information=== | === Additional information=== | ||
| − | |||
=The protein= | =The protein= | ||
| Line 82: | Line 77: | ||
* '''Kinetic information:''' | * '''Kinetic information:''' | ||
| − | * '''Domains:''' | + | * '''[[Domains]]:''' |
* '''Modification:''' | * '''Modification:''' | ||
| − | * ''' | + | * '''[[Cofactors]]:''' |
* '''Effectors of protein activity:''' subject to feedback inhibtion by tryptophan {{PubMed|4956345}} | * '''Effectors of protein activity:''' subject to feedback inhibtion by tryptophan {{PubMed|4956345}} | ||
| Line 113: | Line 108: | ||
* '''Expression browser:''' [http://genome.jouy.inra.fr/cgi-bin/seb/viewdetail.py?id=trpE_2375869_2377416_-1 trpE] {{PubMed|22383849}} | * '''Expression browser:''' [http://genome.jouy.inra.fr/cgi-bin/seb/viewdetail.py?id=trpE_2375869_2377416_-1 trpE] {{PubMed|22383849}} | ||
| − | * '''Sigma factor:''' | + | * '''[[Sigma factor]]:''' |
** ''[[trpE]]'': [[SigA]] {{PubMed|6436812}} | ** ''[[trpE]]'': [[SigA]] {{PubMed|6436812}} | ||
| Line 148: | Line 143: | ||
<pubmed>19385727, ,16285852 </pubmed> | <pubmed>19385727, ,16285852 </pubmed> | ||
==The ''trpE'' [[RNA switch]]== | ==The ''trpE'' [[RNA switch]]== | ||
| − | <pubmed> 20384694 19033375, 17881743, 7515880,2422155,3133360, 7678334,7592410, 14976255, 2422155,8419914, 1551827, 16285852 10714985 11566991 12963367 14712717 21097886 </pubmed> | + | <pubmed> 20384694 19033375, 17881743, 7515880,2422155,3133360, 7678334,7592410, 14976255, 2422155,8419914, 1551827, 16285852 10714985 11566991 12963367 14712717 21097886 24505391 </pubmed> |
==Other original publications== | ==Other original publications== | ||
<pubmed> 4956345 3924737, 6436812, 21815947 </pubmed> | <pubmed> 4956345 3924737, 6436812, 21815947 </pubmed> | ||
[[Category:Protein-coding genes]] | [[Category:Protein-coding genes]] | ||
Revision as of 18:32, 9 February 2014
- Description: anthranilate synthase (subunit I)
| Gene name | trpE |
| Synonyms | |
| Essential | no |
| Product | anthranilate synthase (subunit I) |
| Function | biosynthesis of tryptophan |
| Gene expression levels in SubtiExpress: trpE | |
| Interactions involving this protein in SubtInteract: TrpE | |
| Metabolic function and regulation of this protein in SubtiPathways: trpE | |
| MW, pI | 57 kDa, 5.246 |
| Gene length, protein length | 1545 bp, 515 aa |
| Immediate neighbours | trpD, aroH |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 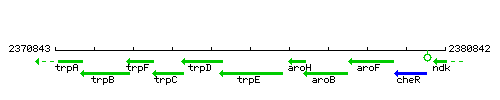
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed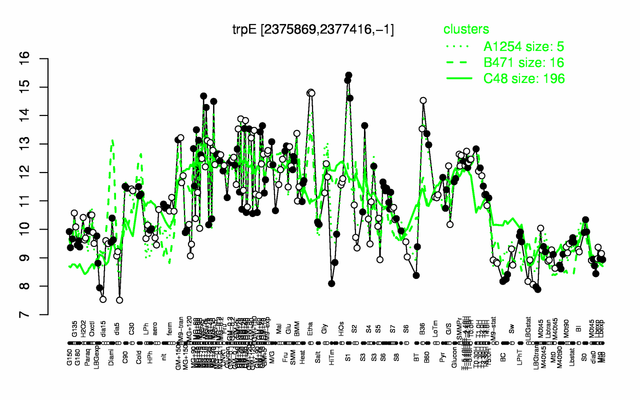
| |
Contents
Categories containing this gene/protein
biosynthesis/ acquisition of amino acids
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU22680
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: Chorismate + L-glutamine = anthranilate + pyruvate + L-glutamate (according to Swiss-Prot)
- Protein family:
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Modification:
- Effectors of protein activity: subject to feedback inhibtion by tryptophan PubMed
Database entries
- UniProt: P03963
- KEGG entry: [3]
- E.C. number: 4.1.3.27
Additional information
Expression and regulation
- Operon:
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Reviews
The trpE RNA switch
Other original publications