Difference between revisions of "LicT"
| Line 33: | Line 33: | ||
<div align="right"> <small>This image was kindly provided by [http://genolist.pasteur.fr/SubtiList/ SubtiList]</small></div> | <div align="right"> <small>This image was kindly provided by [http://genolist.pasteur.fr/SubtiList/ SubtiList]</small></div> | ||
|- | |- | ||
| − | |colspan="2" |'''[http://genome.jouy.inra.fr/cgi-bin/seb/viewdetail.py?id=licT_4012866_4013699_-1 Expression at a glance]'''   {{PubMed|22383849}}<br/>[[Image:licT_expression.png|500px]] | + | |colspan="2" |'''[http://genome.jouy.inra.fr/cgi-bin/seb/viewdetail.py?id=licT_4012866_4013699_-1 Expression at a glance]'''   {{PubMed|22383849}}<br/>[[Image:licT_expression.png|500px|link=http://subtiwiki.uni-goettingen.de/apps/expression/expression.php?search=BSU39080]] |
|- | |- | ||
|} | |} | ||
Revision as of 15:57, 16 May 2013
- Description: transcriptional antiterminator of the bglP-bglH-yxiE operon and the bglS gene
| Gene name | licT |
| Synonyms | |
| Essential | no |
| Product | transcriptional antiterminator (BglG family) |
| Function | control of beta-glucan and beta-glucoside utilization |
| Gene expression levels in SubtiExpress: licT | |
| Interactions involving this protein in SubtInteract: LicT | |
| Metabolic function and regulation of this protein in SubtiPathways: Sugar catabolism | |
| MW, pI | 32 kDa, 5.944 |
| Gene length, protein length | 831 bp, 277 aa |
| Immediate neighbours | bglS, yxiP |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 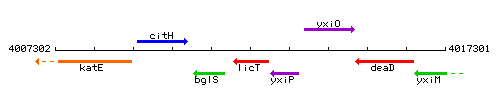
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed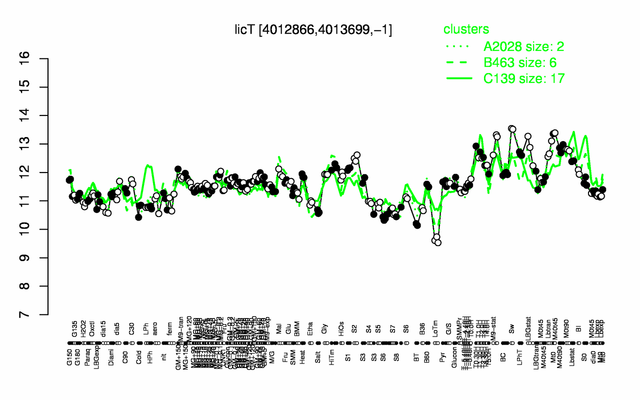
| |
Contents
Categories containing this gene/protein
utilization of specific carbon sources, transcription factors and their control, RNA binding regulators, phosphoproteins
This gene is a member of the following regulons
The LicT regulon: bglP-bglH-yxiE, bglS
The gene
Basic information
- Locus tag: BSU39080
Phenotypes of a mutant
no expression of the bglP-bglH operon
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: binding to the mRNAs of bglS and the bglP-bglH operon, causes transcription antitermination (in presence of salicin and absence of glucose)
- Protein family: transcriptional antiterminator BglG family of antiterminators (according to Swiss-Prot)
Extended information on the protein
- Kinetic information:
- K(D) for the RAT-RNA: 10 nM PubMed
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Localization:
- cytoplasm, even distribution in the absence of the inducer salicin, subpolar localization in the presence of salicin PubMed
Database entries
- UniProt: P39805
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Regulation:
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant: GP427 (licTS, erm), available in the Stülke lab
- Expression vector:
- for expression, purification of both PRDs in E. coli with N-terminal His-tag, in pWH844: pGP165, available in Stülke lab
- for expression, purification of the RNA-binding domain in E. coli with N-terminal His-tag, in pWH844: pGP315, available in Stülke lab
- for expression, purification of the RNA-binding domain in E. coli with N-terminal His-tag and thrombin cleavage site, in pGP570: pGP572, available in Stülke lab
- lacZ fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Stephane Aymerich, Microbiology and Molecular Genetics, INRA Paris-Grignon, France
Josef Deutscher, Microbiology and Molecular Genetics, INRA Paris-Grignon, France
Michael Hecker, Greifswald, Germany Homepage
Your additional remarks
References
Original description
K Schnetz, J Stülke, S Gertz, S Krüger, M Krieg, M Hecker, B Rak
LicT, a Bacillus subtilis transcriptional antiterminator protein of the BglG family.
J Bacteriol: 1996, 178(7);1971-9
[PubMed:8606172]
[WorldCat.org]
[DOI]
(P p)
Control of LicT activity
Structural analysis of LicT
LicT-RNA interaction
Caroline Clerte, Nathalie Declerck, Emmanuel Margeat
Competitive folding of anti-terminator/terminator hairpins monitored by single molecule FRET.
Nucleic Acids Res: 2013, 41(4);2632-43
[PubMed:23303779]
[WorldCat.org]
[DOI]
(I p)
Sebastian Hübner, Nathalie Declerck, Christine Diethmaier, Dominique Le Coq, Stephane Aymerich, Jörg Stülke
Prevention of cross-talk in conserved regulatory systems: identification of specificity determinants in RNA-binding anti-termination proteins of the BglG family.
Nucleic Acids Res: 2011, 39(10);4360-72
[PubMed:21278164]
[WorldCat.org]
[DOI]
(I p)
Hélène Déméné, Thierry Ducat, Karine De Guillen, Catherine Birck, Stéphane Aymerich, Michel Kochoyan, Nathalie Declerck
Structural mechanism of signal transduction between the RNA-binding domain and the phosphotransferase system regulation domain of the LicT antiterminator.
J Biol Chem: 2008, 283(45);30838-49
[PubMed:18682383]
[WorldCat.org]
[DOI]
(P p)
Yinshan Yang, Nathalie Declerck, Xavier Manival, Stéphane Aymerich, Michel Kochoyan
Solution structure of the LicT-RNA antitermination complex: CAT clamping RAT.
EMBO J: 2002, 21(8);1987-97
[PubMed:11953318]
[WorldCat.org]
[DOI]
(P p)
N Declerck, F Vincent, F Hoh, S Aymerich, H van Tilbeurgh
RNA recognition by transcriptional antiterminators of the BglG/SacY family: functional and structural comparison of the CAT domain from SacY and LicT.
J Mol Biol: 1999, 294(2);389-402
[PubMed:10610766]
[WorldCat.org]
[DOI]
(P p)
S Aymerich, M Steinmetz
Specificity determinants and structural features in the RNA target of the bacterial antiterminator proteins of the BglG/SacY family.
Proc Natl Acad Sci U S A: 1992, 89(21);10410-4
[PubMed:1279678]
[WorldCat.org]
[DOI]
(P p)