Difference between revisions of "MtlR"
(→References) |
(→References) |
||
| Line 150: | Line 150: | ||
=References= | =References= | ||
'''Additional publications:''' {{PubMed|23279188,22014119}} | '''Additional publications:''' {{PubMed|23279188,22014119}} | ||
| − | <pubmed>12897001, 20444094,10627040, 10702268 9988713 | + | <pubmed>22900538 </pubmed> |
| − | + | <pubmed>12897001, 20444094,10627040, 10702268 9988713 </pubmed> | |
[[Category:Protein-coding genes]] | [[Category:Protein-coding genes]] | ||
Revision as of 11:16, 3 January 2013
| Gene name | mtlR |
| Synonyms | ydaA |
| Essential | no |
| Product | transcriptional activator, PRD-type |
| Function | regulation of mannitol utilization |
| Gene expression levels in SubtiExpress: mtlR | |
| Interactions involving this protein in SubtInteract: MtlR | |
| Metabolic function and regulation of this protein in SubtiPathways: Sugar catabolism | |
| MW, pI | 78 kDa, 5.313 |
| Gene length, protein length | 2082 bp, 694 aa |
| Immediate neighbours | ycsN, ydaB |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 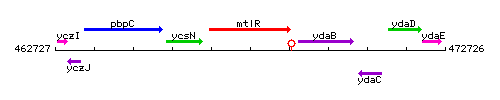
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed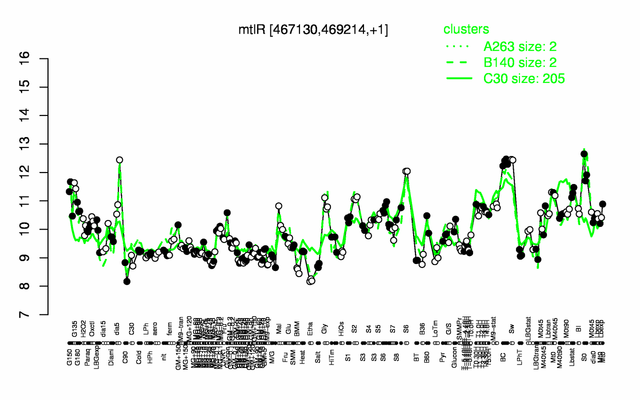
| |
Contents
Categories containing this gene/protein
utilization of specific carbon sources, transcription factors and their control, phosphoproteins
This gene is a member of the following regulons
The MtlR regulon:
The gene
Basic information
- Locus tag: BSU04160
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family:
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains: N-terminal DNA-binding domains, two PTS-regulation domains (PRD1 and PRD2), EIIB (Gat)-like domain, EIIA (Mtl)-like domain
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- activity is stimulated by PtsH-dependent phosphorylation in PRD2 (mechnism of carbon catabolite repression) PubMed
- activity is inhibited by MtlF-dependent phosphorylation in the EIIB(Gat)-like domain on Cys-419 (this prevents activity in the absence of mannitol and allows induction in presence of mannitol) PubMed
Database entries
- Structure:
- UniProt: P96574
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Operon: mtlR PubMed
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Additional publications: PubMed