Difference between revisions of "LicT"
| Line 34: | Line 34: | ||
<br/><br/><br/><br/><br/><br/> | <br/><br/><br/><br/><br/><br/> | ||
| + | |||
| + | = [[Categories]] containing this gene/protein = | ||
| + | {{SubtiWiki category|[[utilization of specific carbon sources]]}}, | ||
| + | {{SubtiWiki category|[[transcription factors and their control]]}}, | ||
| + | {{SubtiWiki category|[[RNA binding regulators]]}}, | ||
| + | {{SubtiWiki category|[[phosphoproteins]]}} | ||
| + | |||
| + | = This gene is a member of the following [[regulons]] = | ||
| + | |||
=The gene= | =The gene= | ||
| Line 55: | Line 64: | ||
| − | + | ||
| − | |||
| − | |||
| − | |||
| − | |||
=The protein= | =The protein= | ||
Revision as of 01:54, 9 December 2010
- Description: transcriptional antiterminator of the bglP-bglH operon and the bglS gene
| Gene name | licT |
| Synonyms | |
| Essential | no |
| Product | transcriptional antiterminator (BglG family) |
| Function | substrate-dependent induction of bglP-bglH and bglS |
| Metabolic function and regulation of this protein in SubtiPathways: Sugar catabolism | |
| MW, pI | 32 kDa, 5.944 |
| Gene length, protein length | 831 bp, 277 aa |
| Immediate neighbours | bglS, yxiP |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 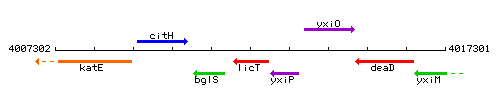
This image was kindly provided by SubtiList
| |
Contents
Categories containing this gene/protein
utilization of specific carbon sources, transcription factors and their control, RNA binding regulators, phosphoproteins
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU39080
Phenotypes of a mutant
no expression of the bglP-bglH operon
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: binding to the mRNAs of bglS and the bglP-bglH operon, causes transcription antitermination (in presence of salicin and absence of glucose)
- Protein family: transcriptional antiterminator BglG family of antiterminators (according to Swiss-Prot)
Extended information on the protein
- Kinetic information:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Localization:
Database entries
- UniProt: P39805
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Regulation:
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant: GP427 (licTS, erm), available in the Stülke lab
- Expression vector:
- for expression, purification of both PRDs in E. coli with N-terminal His-tag, in pWH844: pGP165, available in Stülke lab
- for expression, purification of the RNA-binding domain in E. coli with N-terminal His-tag, in pWH844: pGP315, available in Stülke lab
- for expression, purification of the RNA-binding domain in E. coli with N-terminal His-tag and thrombin cleavage site, in pGP570: pGP572, available in Stülke lab
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Stephane Aymerich, Microbiology and Molecular Genetics, INRA Paris-Grignon, France
Josef Deutscher, Microbiology and Molecular Genetics, INRA Paris-Grignon, France
Michael Hecker, Greifswald, Germany Homepage
Your additional remarks
References
Original description
Control of LicT activity
Structural analysis of LicT
LicT-RNA interaction