Difference between revisions of "AbrB"
| Line 64: | Line 64: | ||
=== Genes/ operons controlled by AbrB === | === Genes/ operons controlled by AbrB === | ||
| − | * '''Activated by AbrB:''' ''[[citB]]'', '' | + | * '''Activated by AbrB:''' ''[[citB]]'', ''[[hpr]]'', ''[[rbsR]]-[[rbsK]]-[[rbsD]]-[[rbsA]]-[[rbsC]]-[[rbsB]]'' |
| − | * ''' Repressed by AbrB:''' ''[[abrB]], [[aprE]], [[ftsA]]-[[ftsZ]], [[kinC]], [[motA]], [[nprE]], [[pbpE]], [[spo0H]], [[spoVG]], [[spo0E]], [[tycA]], [[sbo]]-[[alb]], [[yqxM]]-[[sipW]]-[[tasA]], [[sdpA operon]], [[sunA operon]], [[skfA operon]], [[abh]], [[pbpE]], [[ggt]], [[sinI]]-[[sinR]], [[sigW]], [[yuaB]]'' | + | * ''' Repressed by AbrB:''' ''[[abrB]], [[aprE]], [[comK]], [[ftsA]]-[[ftsZ]], [[kinC]], [[motA]], [[nprE]], [[pbpE]], [[spo0H]], [[spoVG]], [[spo0E]], [[tycA]], [[sbo]]-[[alb]], [[yqxM]]-[[sipW]]-[[tasA]], [[sdpA operon]], [[sunA operon]], [[skfA operon]], [[abh]], [[pbpE]], [[ggt]], [[sinI]]-[[sinR]], [[sigW]], [[yuaB]]'' |
=== Extended information on the protein === | === Extended information on the protein === | ||
Revision as of 15:38, 22 April 2010
- Description: transcriptional regulator of transition state genes
| Gene name | abrB |
| Synonyms | cpsX, tolB |
| Essential | no |
| Product | transcriptional regulator |
| Function | regulation of gene expression during the transition from growth to stationary phase |
| Metabolic function and regulation of this protein in SubtiPathways: Biofilm, His, Asp, Asn, Ammonium/ glutamate, Central C-metabolism, Sugar catabolism | |
| MW, pI | 10 kDa, 6.57 |
| Gene length, protein length | 282 bp, 94 aa |
| Immediate neighbours | yabC, metS |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 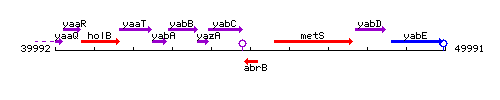
This image was kindly provided by SubtiList
| |
Contents
The gene
Basic information
- Locus tag: BSU00370
Phenotypes of a mutant
No swarming motility on B medium. PubMed
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family:
Genes/ operons controlled by AbrB
- Repressed by AbrB: abrB, aprE, comK, ftsA-ftsZ, kinC, motA, nprE, pbpE, spo0H, spoVG, spo0E, tycA, sbo-alb, yqxM-sipW-tasA, sdpA operon, sunA operon, skfA operon, abh, pbpE, ggt, sinI-sinR, sigW, yuaB
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Localization:
Database entries
- UniProt: P08874
- KEGG entry: [3]
Additional information
Expression and regulation
- Operon: abrB PubMed
- Regulation: expressed at the onset of stationary phase PubMed
- Additional information:
Biological materials
- Mutant: TT731 (aphA3)
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Richard Losick, Harvard Univ., Cambridge, USA homepage
Mark Strauch, Baltimore, USA homepage
Your additional remarks
References
Reviews
Original Publications