Difference between revisions of "ClpP"
(→Original Publications) |
|||
| Line 149: | Line 149: | ||
* '''Mutant:''' | * '''Mutant:''' | ||
** ''clpP::spec'' and ''clpP::cat'', available in the [[Leendert Hamoen]] lab | ** ''clpP::spec'' and ''clpP::cat'', available in the [[Leendert Hamoen]] lab | ||
| − | ** GP551 (spc), available in | + | ** BP99 (''clpP''::''tet''), available in [[Fabian Commichau]]'s lab {{PubMed|25610436}} |
| − | ** 1S139 ( ''clpP''::''erm''), {{PubMed| }}, available at [http://pasture.asc.ohio-state.edu/BGSC/getdetail.cfm?bgscid=1S139&Search=1S139 BGSC] | + | ** GP551 (spc), available in [[Jörg Stülke]]'s lab |
| + | ** 1S139 (''clpP''::''erm''), {{PubMed| }}, available at [http://pasture.asc.ohio-state.edu/BGSC/getdetail.cfm?bgscid=1S139&Search=1S139 BGSC] | ||
** 1S140 ( ''clpP''::''spec''), {{PubMed| }}, available at [http://pasture.asc.ohio-state.edu/BGSC/getdetail.cfm?bgscid=1S140&Search=1S140 BGSC] | ** 1S140 ( ''clpP''::''spec''), {{PubMed| }}, available at [http://pasture.asc.ohio-state.edu/BGSC/getdetail.cfm?bgscid=1S140&Search=1S140 BGSC] | ||
| Line 164: | Line 165: | ||
=Labs working on this gene/protein= | =Labs working on this gene/protein= | ||
| − | [[Leendert Hamoen]], Newcastle University, UK [http://www.ncl.ac.uk/camb/staff/profile/l.hamoen homepage] | + | * [[Leendert Hamoen]], Newcastle University, UK [http://www.ncl.ac.uk/camb/staff/profile/l.hamoen homepage] |
=Your additional remarks= | =Your additional remarks= | ||
| Line 172: | Line 173: | ||
<pubmed> 16211032 17302811 23375660 23479438,19609260,19781636</pubmed> | <pubmed> 16211032 17302811 23375660 23479438,19609260,19781636</pubmed> | ||
==Original Publications== | ==Original Publications== | ||
| − | <pubmed>9643546,10809708,11807061,9987115,14679237,18689476,15317791,17586624,11684022,12923101,17560370, 16899079,19226326 , 20070525,9987115,11544224 14763982 9643546, 19767395 11112444 9535081 18689473 20305655 20852588 16200071 21969594 22080375 22517742 17380125,12598648,9890793,20049702,20049702 23927726 24263382 24226776 24417481 15378759 23361916 24942655 25212124 25433860 </pubmed> | + | <pubmed>9643546,10809708,11807061,9987115,14679237,18689476,15317791,17586624,11684022,12923101,17560370, 16899079,19226326 , 20070525,9987115,11544224 14763982 9643546, 19767395 11112444 9535081 18689473 20305655 20852588 16200071 21969594 22080375 22517742 17380125,12598648,9890793,20049702,20049702 23927726 24263382 24226776 24417481 15378759 23361916 24942655 25212124 25433860 25610436</pubmed> |
[[Category:Protein-coding genes]] | [[Category:Protein-coding genes]] | ||
Revision as of 10:04, 23 January 2015
- Description: ATP-dependent Clp protease proteolytic subunit (class III heat-shock protein)
| Gene name | clpP |
| Synonyms | yvdN |
| Essential | no |
| Product | ATP-dependent Clp protease proteolytic subunit |
| Function | protein degradation |
| Gene expression levels in SubtiExpress: clpP | |
| Interactions involving this protein in SubtInteract: ClpP | |
| Metabolic function and regulation of this protein in SubtiPathways: clpP | |
| MW, pI | 21 kDa, 5.008 |
| Gene length, protein length | 591 bp, 197 aa |
| Immediate neighbours | trnQ-Arg, pgcM |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 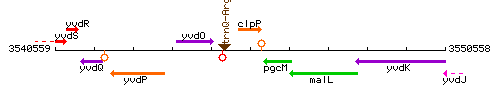
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed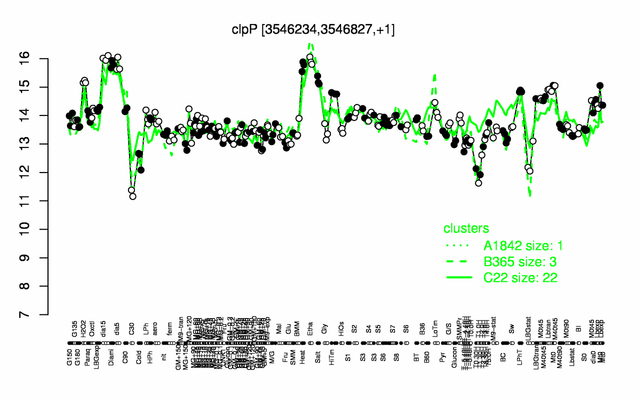
| |
Contents
Categories containing this gene/protein
proteolysis, general stress proteins (controlled by SigB), heat shock proteins, phosphoproteins, most abundant proteins
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU34540
Phenotypes of a mutant
- increased thermotolerance due to increased stabiliy of Spx and thus increased expression of trxA PubMed
Database entries
- BsubCyc: BSU34540
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: Hydrolysis of proteins to small peptides in the presence of ATP and magnesium (according to Swiss-Prot) endopeptidase/proteolysis
- Protein family: peptidase S14 family (according to Swiss-Prot) ClpP (IPR001907) InterPro, (PF00574) PFAM
- Paralogous protein(s):
Targets of ClpC-ClpP-dependent protein degradation
Targets of ClpX-ClpP-dependent protein degradation
Extended information on the protein
- Kinetic information:
- Modification:
- phosphorylated on Arg-13 PubMed
- Effectors of protein activity:
Database entries
- BsubCyc: BSU34540
- UniProt: P80244
- KEGG entry: [3]
- E.C. number: 3.4.21.92
Additional information
Expression and regulation
- Operon: clpP PubMed
- Additional information:
- belongs to the 100 most abundant proteins PubMed
- number of protein molecules per cell (minimal medium with glucose and ammonium): 4011 PubMed
- number of protein molecules per cell (complex medium with amino acids, without glucose): 11118 PubMed
- number of protein molecules per cell (minimal medium with glucose and ammonium, exponential phase): 1536 PubMed
- number of protein molecules per cell (minimal medium with glucose and ammonium, early stationary phase after glucose exhaustion): 731 PubMed
- number of protein molecules per cell (minimal medium with glucose and ammonium, late stationary phase after glucose exhaustion): 1056 PubMed
Biological materials
- Mutant:
- clpP::spec and clpP::cat, available in the Leendert Hamoen lab
- BP99 (clpP::tet), available in Fabian Commichau's lab PubMed
- GP551 (spc), available in Jörg Stülke's lab
- 1S139 (clpP::erm), PubMed, available at BGSC
- 1S140 ( clpP::spec), PubMed, available at BGSC
- Expression vector:
- lacZ fusion:
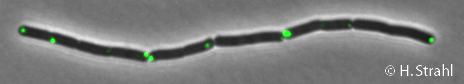
- GFP fusion: C-terminal GFP fusions (both single copy and as 2th copy in amyE locus, also as CFP and YFP variants) available in the Leendert Hamoen lab
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
- Leendert Hamoen, Newcastle University, UK homepage
Your additional remarks
References
Reviews
Original Publications