Difference between revisions of "LicC"
| Line 49: | Line 49: | ||
{{SubtiWiki category|[[utilization of specific carbon sources]]}}, | {{SubtiWiki category|[[utilization of specific carbon sources]]}}, | ||
{{SubtiWiki category|[[membrane proteins]]}}, | {{SubtiWiki category|[[membrane proteins]]}}, | ||
| − | {{SubtiWiki category|[[sporulation | + | {{SubtiWiki category|[[Sporulation#Sporulation_proteins/_other|sporulation proteins]]}} |
= This gene is a member of the following [[regulons]] = | = This gene is a member of the following [[regulons]] = | ||
Revision as of 12:04, 24 April 2014
- Description: lichenan-specific phosphotransferase system, EIIC component of the PTS
| Gene name | licC |
| Synonyms | celB |
| Essential | no |
| Product | lichenan-specific phosphotransferase system, EIIC component |
| Function | lichenan uptake and phosphorylation |
| Gene expression levels in SubtiExpress: licC | |
| Interactions involving this protein in SubtInteract: LicC | |
| Metabolic function and regulation of this protein in SubtiPathways: licC | |
| MW, pI | 48 kDa, 8.584 |
| Gene length, protein length | 1356 bp, 452 aa |
| Immediate neighbours | licA, licB |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 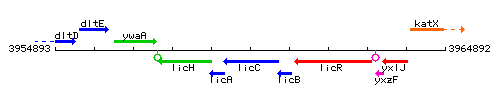
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed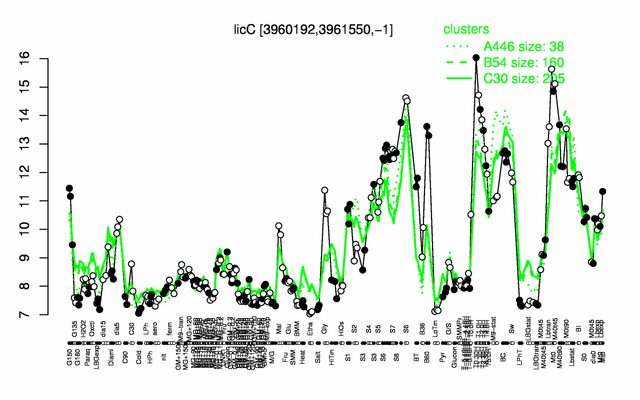
| |
Contents
Categories containing this gene/protein
phosphotransferase systems, utilization of specific carbon sources, membrane proteins, sporulation proteins
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU38580
Phenotypes of a mutant
Database entries
- BsubCyc: BSU38580
- DBTBS entry: no entry
- SubtiList entry: [1]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Localization: cell membrane (according to Swiss-Prot)
Database entries
- BsubCyc: BSU38580
- Structure: 3QNQ (EIIC component of diacetylchitobiose specific PTS from Bacillus cereus; 60% identity, 87% similarity) PubMed
- UniProt: P46317
- KEGG entry: [2]
- E.C. number:
Additional information
Expression and regulation
- Regulation:
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Additional publications: PubMed