Difference between revisions of "PdhD"
| Line 16: | Line 16: | ||
|colspan="2" style="background:#FAF8CC;" align="center"| '''Gene expression levels in [http://subtiwiki.uni-goettingen.de/apps/expression/ ''Subti''Express]''': [http://subtiwiki.uni-goettingen.de/apps/expression/expression.php?search=BSU14610 pdhD] | |colspan="2" style="background:#FAF8CC;" align="center"| '''Gene expression levels in [http://subtiwiki.uni-goettingen.de/apps/expression/ ''Subti''Express]''': [http://subtiwiki.uni-goettingen.de/apps/expression/expression.php?search=BSU14610 pdhD] | ||
|- | |- | ||
| − | |colspan="2" style="background:#FAF8CC;" align="center"| '''Interactions involving this protein in [http:// | + | |colspan="2" style="background:#FAF8CC;" align="center"| '''Interactions involving this protein in [http://subtiwiki.uni-goettingen.de/interact/ ''Subt''Interact]''': [http://subtiwiki.uni-goettingen.de/interact/index.php?protein=PdhD PdhD] |
|- | |- | ||
|colspan="2" style="background:#FAF8CC;" align="center"| '''Metabolic function and regulation of this protein in [[SubtiPathways|''Subti''Pathways]]: <br/>[http://subtiwiki.uni-goettingen.de/pathways/carbon_flow.html Central C-metabolism]''' | |colspan="2" style="background:#FAF8CC;" align="center"| '''Metabolic function and regulation of this protein in [[SubtiPathways|''Subti''Pathways]]: <br/>[http://subtiwiki.uni-goettingen.de/pathways/carbon_flow.html Central C-metabolism]''' | ||
Revision as of 17:10, 11 November 2013
- Description: dihydrolipoamide dehydrogenase E3 subunit of both pyruvate dehydrogenase and 2-oxoglutarate dehydrogenase complexes
| Gene name | pdhD |
| Synonyms | citL |
| Essential | no |
| Product | dihydrolipoamide dehydrogenase E3 subunit of both pyruvate dehydrogenase and 2-oxoglutarate dehydrogenase complexes |
| Function | links glycolysis and TCA cycle, enzyme in TCA cycle |
| Gene expression levels in SubtiExpress: pdhD | |
| Interactions involving this protein in SubtInteract: PdhD | |
| Metabolic function and regulation of this protein in SubtiPathways: Central C-metabolism | |
| MW, pI | 49 kDa, 4.76 |
| Gene length, protein length | 1410 bp, 470 aa |
| Immediate neighbours | pdhC, slp |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 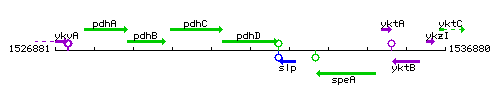
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed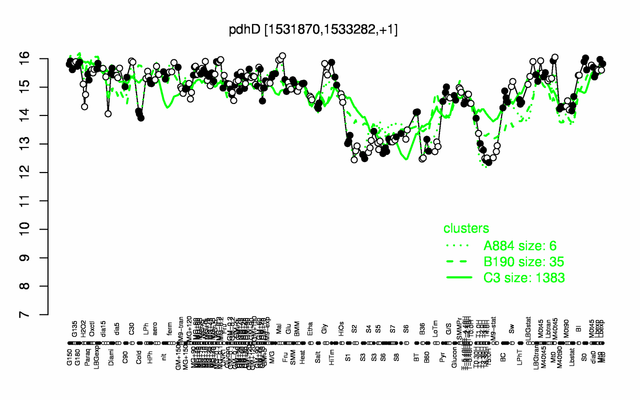
| |
Contents
Categories containing this gene/protein
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU14610
Phenotypes of a mutant
- defects in sporulation and unable to grow on glucose as single carbon source PubMed
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: Protein N(6)-(dihydrolipoyl)lysine + NAD+ = protein N(6)-(lipoyl)lysine + NADH (according to Swiss-Prot)
- Protein family: class-I pyridine nucleotide-disulfide oxidoreductase family (according to Swiss-Prot)
Extended information on the protein
- Kinetic information: Michaelis-Menten PubMed
- Domains:
- Modification: phosphorylated (Ser/Thr/Tyr) PubMed
- Cofactor(s):
- Effectors of protein activity:
- Localization: cytoplasm (according to Swiss-Prot)
Database entries
- Structure: 1EBD (complex with binding domain of dihydrolipoamide acetylase, Geobacillus stearothermophilus), 1EBD (complex with binding domain of dihydrolipoamide acetylase, Geobacillus stearothermophilus)
- UniProt: P21880
- KEGG entry: [3]
- E.C. number: 1.8.1.4
Additional information
Expression and regulation
- Regulation:
- Regulatory mechanism:
- stringent response: due to presence of guanine at +1 position of the transcript PubMed
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- GFP fusion:
- two-hybrid system: B. pertussis adenylate cyclase-based bacterial two hybrid system (BACTH), available in Stülke lab
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Reviews
Kai Tittmann
Reaction mechanisms of thiamin diphosphate enzymes: redox reactions.
FEBS J: 2009, 276(9);2454-68
[PubMed:19476487]
[WorldCat.org]
[DOI]
(I p)
K F Sheu, J P Blass
The alpha-ketoglutarate dehydrogenase complex.
Ann N Y Acad Sci: 1999, 893;61-78
[PubMed:10672230]
[WorldCat.org]
[DOI]
(P p)
U Neveling, S Bringer-Meyer, H Sahm
Gene and subunit organization of bacterial pyruvate dehydrogenase complexes.
Biochim Biophys Acta: 1998, 1385(2);367-72
[PubMed:9655937]
[WorldCat.org]
[DOI]
(P p)
M S Patel, T E Roche
Molecular biology and biochemistry of pyruvate dehydrogenase complexes.
FASEB J: 1990, 4(14);3224-33
[PubMed:2227213]
[WorldCat.org]
[DOI]
(P p)
P A Frey
Mechanism of coupled electron and group transfer in Escherichia coli pyruvate dehydrogenase.
Ann N Y Acad Sci: 1982, 378;250-64
[PubMed:6805383]
[WorldCat.org]
[DOI]
(P p)
Original publications