Difference between revisions of "CwlO"
(→References) |
|||
| Line 141: | Line 141: | ||
=References= | =References= | ||
| + | == Reviews == | ||
| + | <pubmed> 23066944 </pubmed> | ||
| + | == Original publications == | ||
'''Additional publications:''' {{PubMed|21478646}} | '''Additional publications:''' {{PubMed|21478646}} | ||
<pubmed>16233686,17581128, 20525796,18957862, 20059685 ,22139507</pubmed> | <pubmed>16233686,17581128, 20525796,18957862, 20059685 ,22139507</pubmed> | ||
[[Category:Protein-coding genes]] | [[Category:Protein-coding genes]] | ||
Revision as of 10:29, 17 October 2012
- Description: D,L-endopeptidase-type autolysin
| Gene name | cwlO |
| Synonyms | yzkA, yvcE |
| Essential | no |
| Product | endopeptidase-type autolysin |
| Function | cell wall synthesis, cell proliferation |
| Gene expression levels in SubtiExpress: cwlO | |
| MW, pI | 50 kDa, 5.326 |
| Gene length, protein length | 1419 bp, 473 aa |
| Immediate neighbours | trxB, yvcD |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 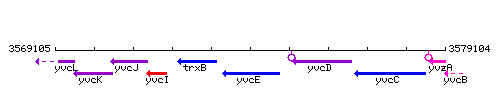
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed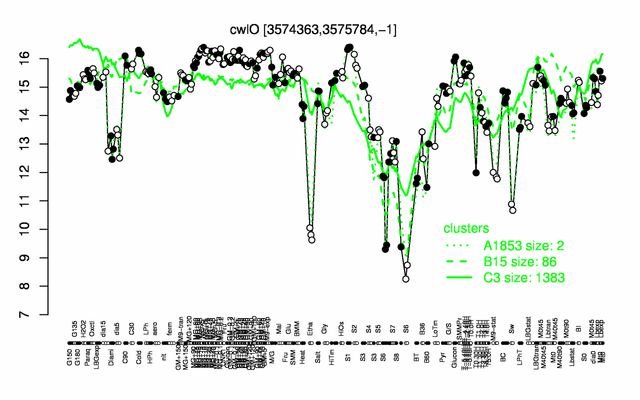
| |
Contents
Categories containing this gene/protein
cell wall degradation/ turnover
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU34800
Phenotypes of a mutant
a cwlO lytE mutant is not viable PubMed
Database entries
- DBTBS entry: no entry
- SubtiList entry: [1]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family: peptidase C40 family (according to Swiss-Prot)
- Paralogous protein(s): the C-terminal D,L-endopeptidase domains of LytE, LytF, CwlS, and CwlO exhibit strong sequence similarity
Extended information on the protein
- Kinetic information:
- Domains:
- C-terminal D,L-endopeptidase domain PubMed
- Modification:
- Cofactor(s):
- Effectors of protein activity:
Database entries
- Structure:
- UniProt: P40767
- KEGG entry: [2]
- E.C. number:
Additional information
Expression and regulation
- Operon:
- Regulatory mechanism:
- Additional information:
- The mRNA has a long 5' leader region. This may indicate RNA-based regulation PubMed
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Reviews
Waldemar Vollmer
Bacterial growth does require peptidoglycan hydrolases.
Mol Microbiol: 2012, 86(5);1031-5
[PubMed:23066944]
[WorldCat.org]
[DOI]
(I p)
Original publications
Additional publications: PubMed