Difference between revisions of "MtnX"
| Line 26: | Line 26: | ||
|colspan="2" | '''Genetic context''' <br/> [[Image:ykrX_context.gif]] | |colspan="2" | '''Genetic context''' <br/> [[Image:ykrX_context.gif]] | ||
<div align="right"> <small>This image was kindly provided by [http://genolist.pasteur.fr/SubtiList/ SubtiList]</small></div> | <div align="right"> <small>This image was kindly provided by [http://genolist.pasteur.fr/SubtiList/ SubtiList]</small></div> | ||
| + | |- | ||
| + | |colspan="2" |'''[http://genome.jouy.inra.fr/cgi-bin/seb/viewdetail.py?id=mtnX_1428275_1428982_1 Expression at a glance]'''   {{PubMed|22383849}}<br/>[[Image:mtnX_expression.png|500px]] | ||
|- | |- | ||
|} | |} | ||
__TOC__ | __TOC__ | ||
| + | <br/><br/><br/><br/> | ||
| + | <br/><br/><br/><br/> | ||
| + | <br/><br/><br/><br/> | ||
| + | <br/><br/><br/><br/> | ||
| + | <br/><br/><br/><br/> | ||
| + | |||
<br/><br/><br/><br/><br/><br/> | <br/><br/><br/><br/><br/><br/> | ||
Revision as of 09:12, 19 April 2012
- Description: 2-hydroxy-3-keto-5-methylthiopentenyl-1-phosphate phosphatase
| Gene name | mtnX |
| Synonyms | ykrX |
| Essential | no |
| Product | 2-hydroxy-3-keto-5-methylthiopentenyl- 1-phosphate phosphatase |
| Function | methionine salvage |
| Metabolic function and regulation of this protein in SubtiPathways: Cys, Met & Sulfate assimilation | |
| MW, pI | 26 kDa, 4.658 |
| Gene length, protein length | 705 bp, 235 aa |
| Immediate neighbours | mtnW, mtnB |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 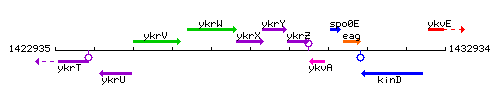
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed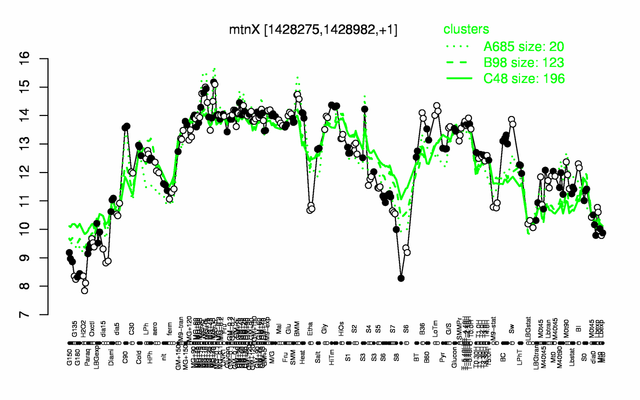
| |
Contents
Categories containing this gene/protein
biosynthesis/ acquisition of amino acids
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU13600
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: 2-hydroxy-5-(methylthio)-3-oxopent-1-enyl phosphate + H2O = 1,2-dihydroxy-5-(methylthio)pent-1-en-3-one + phosphate (according to Swiss-Prot)
- Protein family: MtnX family (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
Database entries
- Structure: 2FEA
- UniProt: O31667
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Regulatory mechanism: S-box: transcription termination/ antitermination, the S-box riboswitch binds S-adenosylmethionine resulting in termination PubMed
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References