Difference between revisions of "CshA"
| Line 26: | Line 26: | ||
|colspan="2" | '''Genetic context''' <br/> [[Image:ydbR_context.gif]] | |colspan="2" | '''Genetic context''' <br/> [[Image:ydbR_context.gif]] | ||
<div align="right"> <small>This image was kindly provided by [http://genolist.pasteur.fr/SubtiList/ SubtiList]</small></div> | <div align="right"> <small>This image was kindly provided by [http://genolist.pasteur.fr/SubtiList/ SubtiList]</small></div> | ||
| + | |- | ||
| + | |colspan="2" |'''[http://genome.jouy.inra.fr/cgi-bin/seb/viewdetail.py?id=cshA_511106_512641_1 Expression at a glance]'''   {{PubMed|22383849}}<br/>[[Image:cshA_expression.png|500px]] | ||
|- | |- | ||
|} | |} | ||
__TOC__ | __TOC__ | ||
| + | <br/><br/><br/><br/> | ||
| + | <br/><br/><br/><br/> | ||
| + | <br/><br/><br/><br/> | ||
| + | <br/><br/><br/><br/> | ||
| + | <br/><br/><br/><br/> | ||
| + | |||
<br/><br/> | <br/><br/> | ||
Revision as of 15:40, 18 April 2012
- Description: DEAD-box RNA helicase, important for adaptation to low temperatures
| Gene name | cshA |
| Synonyms | ydbR |
| Essential | no |
| Product | DEAD-box RNA helicase |
| Function | RNA helicase |
| Interactions involving this protein in SubtInteract: CshA | |
| MW, pI | 57 kDa, 9.89 |
| Gene length, protein length | 1533 bp, 511 aa |
| Immediate neighbours | murF, ydbS |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 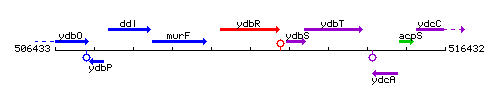
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed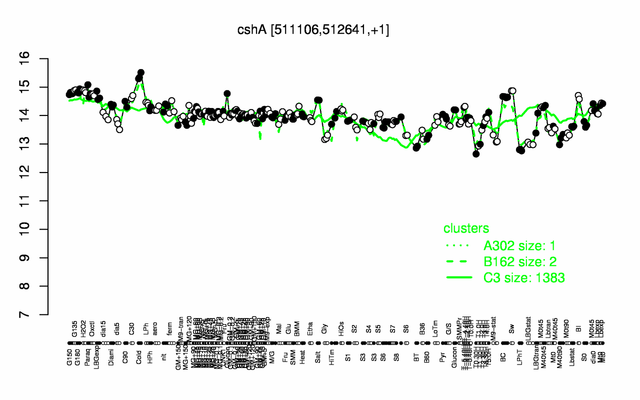
| |
Contents
Categories containing this gene/protein
DEAD-box RNA helicases, cold stress proteins, membrane proteins
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU04580
Phenotypes of a mutant
Database entries
- DBTBS entry: no entry
- SubtiList entry: [1]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: RNA helicase
- Protein family: helicase C-terminal domain (according to Swiss-Prot) DEAD-box RNA helicase
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Localization: cytoplasma, colocalizes with the ribosomes PubMed, cell membrane PubMed
Database entries
- Structure:
- UniProt: P96614
- KEGG entry: [2]
- E.C. number:
Additional information
Expression and regulation
- Operon:
- Sigma factor:
- Regulation:
- Regulatory mechanism:
- Additional information:
Biological materials
- Expression vector:
- for expression/ purification from B. subtilis with C-terminal Strep-tag, for SPINE, in pGP382: pGP1387, available in Stülke lab
- for expression/ purification from B. subtilis with C-terminal Strep-tag, for SPINE, expression from the native chromomsomal site: GP1026, available in Stülke lab
- for expression/ purification from E. coli with N-terminal His-tag, in pWH844: pGP1386, available in Stülke lab
- lacZ fusion:
- GFP fusion: pGP1369 for chromosomal expression of CshA-YFP, available in Stülke lab
- B. subtilis GP1081 cshA-gfp spc, available in Stülke lab,
- two-hybrid system: B. pertussis adenylate cyclase-based bacterial two hybrid system (BACTH), available in Stülke lab
- Antibody:
Labs working on this gene/protein
Mohamed Marahiel, Marburg University, Germany homepage
Your additional remarks
References
Additional publications: PubMed
Martin Lehnik-Habrink, Joseph Newman, Fabian M Rothe, Alexandra S Solovyova, Cecilia Rodrigues, Christina Herzberg, Fabian M Commichau, Richard J Lewis, Jörg Stülke
RNase Y in Bacillus subtilis: a Natively disordered protein that is the functional equivalent of RNase E from Escherichia coli.
J Bacteriol: 2011, 193(19);5431-41
[PubMed:21803996]
[WorldCat.org]
[DOI]
(I p)
Christelle M Roux, Jonathon P DeMuth, Paul M Dunman
Characterization of components of the Staphylococcus aureus mRNA degradosome holoenzyme-like complex.
J Bacteriol: 2011, 193(19);5520-6
[PubMed:21764917]
[WorldCat.org]
[DOI]
(I p)
Martin Lehnik-Habrink, Henrike Pförtner, Leonie Rempeters, Nico Pietack, Christina Herzberg, Jörg Stülke
The RNA degradosome in Bacillus subtilis: identification of CshA as the major RNA helicase in the multiprotein complex.
Mol Microbiol: 2010, 77(4);958-71
[PubMed:20572937]
[WorldCat.org]
[DOI]
(I p)
Yoshinari Ando, Kouji Nakamura
Bacillus subtilis DEAD protein YdbR possesses ATPase, RNA binding, and RNA unwinding activities.
Biosci Biotechnol Biochem: 2006, 70(7);1606-15
[PubMed:16861794]
[WorldCat.org]
[DOI]
(P p)
Karen Hunger, Carsten L Beckering, Frank Wiegeshoff, Peter L Graumann, Mohamed A Marahiel
Cold-induced putative DEAD box RNA helicases CshA and CshB are essential for cold adaptation and interact with cold shock protein B in Bacillus subtilis.
J Bacteriol: 2006, 188(1);240-8
[PubMed:16352840]
[WorldCat.org]
[DOI]
(P p)
Carsten L Beckering, Leif Steil, Michael H W Weber, Uwe Völker, Mohamed A Marahiel
Genomewide transcriptional analysis of the cold shock response in Bacillus subtilis.
J Bacteriol: 2002, 184(22);6395-402
[PubMed:12399512]
[WorldCat.org]
[DOI]
(P p)