Difference between revisions of "RpoB"
(→References) |
|||
| Line 102: | Line 102: | ||
* '''Regulatory mechanism:''' | * '''Regulatory mechanism:''' | ||
| − | * '''Additional information:''' subject to Clp-dependent proteolysis upon glucose starvation [http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=+17981983 PubMed] | + | * '''Additional information:''' |
| + | ** subject to Clp-dependent proteolysis upon glucose starvation [http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=+17981983 PubMed] | ||
| + | ** The mRNA has a long 5' leader region. This may indicate RNA-based regulation {{PubMed|20525796}} | ||
=Biological materials = | =Biological materials = | ||
| Line 124: | Line 126: | ||
=References= | =References= | ||
| − | <pubmed> 19680289 17644585 7657605,12486038,10675340,7592585,, 17981983, 18763711, 19191541 </pubmed> | + | <pubmed> 19680289 17644585 7657605,12486038,10675340,7592585, 20525796, 17981983, 18763711, 19191541 </pubmed> |
[[Category:Protein-coding genes]] | [[Category:Protein-coding genes]] | ||
Revision as of 19:50, 10 June 2010
- Description: RNA polymerase beta subunit
| Gene name | rpoB |
| Synonyms | |
| Essential | yes PubMed |
| Product | RNA polymerase beta subunit |
| Function | transcription |
| MW, pI | 133 kDa, 4.731 |
| Gene length, protein length | 3579 bp, 1193 aa |
| Immediate neighbours | ybxB, rpoC |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 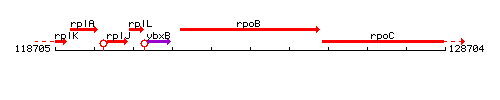
This image was kindly provided by SubtiList
| |
Contents
The gene
Basic information
- Locus tag: BSU01070
Phenotypes of a mutant
essential PubMed
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: Nucleoside triphosphate + RNA(n) = diphosphate + RNA(n+1) (according to Swiss-Prot)
- Protein family: RNA polymerase beta chain family (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Localization: membrane associated PubMed
Database entries
- Structure:
- UniProt: P37870
- KEGG entry: [3]
- E.C. number: 2.7.7.6
Additional information
- subject to Clp-dependent proteolysis upon glucose starvation PubMed
Expression and regulation
- Operon: rpoB DBTBS
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References