Difference between revisions of "CmoO"
| Line 119: | Line 119: | ||
<pubmed>16109943,, </pubmed> | <pubmed>16109943,, </pubmed> | ||
| − | |||
Revision as of 23:13, 15 June 2009
- Description: similar to monooxygenase
| Gene name | ytmO |
| Synonyms | |
| Essential | no |
| Product | unknown |
| Function | unknown |
| MW, pI | 36 kDa, 5.845 |
| Gene length, protein length | 1002 bp, 334 aa |
| Immediate neighbours | ytnI, tcyN |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 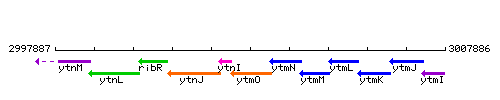
This image was kindly provided by SubtiList
| |
Contents
The gene
Basic information
- Locus tag: BSU29330
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family:
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Interactions:
- Localization:
Database entries
- Structure:
- Swiss prot entry: O34846
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Regulation:
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Pierre Burguière, Juliette Fert, Isabelle Guillouard, Sandrine Auger, Antoine Danchin, Isabelle Martin-Verstraete
Regulation of the Bacillus subtilis ytmI operon, involved in sulfur metabolism.
J Bacteriol: 2005, 187(17);6019-30
[PubMed:16109943]
[WorldCat.org]
[DOI]
(P p)