Difference between revisions of "GmuC"
(→Expression and regulation) |
|||
| Line 90: | Line 90: | ||
* '''Operon:''' ''[[gmuB]]-[[gmuA]]-[[gmuC]]-[[gmuD]]-[[gmuR]]-[[gmuE]]-[[gmuF]]-[[gmuG]]'' [http://www.ncbi.nlm.nih.gov/sites/entrez/18177310 PubMed] | * '''Operon:''' ''[[gmuB]]-[[gmuA]]-[[gmuC]]-[[gmuD]]-[[gmuR]]-[[gmuE]]-[[gmuF]]-[[gmuG]]'' [http://www.ncbi.nlm.nih.gov/sites/entrez/18177310 PubMed] | ||
| − | * '''[[Sigma factor]]:''' [[SigA]] | + | * '''[[Sigma factor]]:''' [[SigA]] [http://www.ncbi.nlm.nih.gov/sites/entrez/18177310 PubMed] |
* '''Regulation:''' | * '''Regulation:''' | ||
| + | ** induced by conjac glucomannan (requires [[GmuG]]) ([[GmuR]]) [http://www.ncbi.nlm.nih.gov/sites/entrez/18177310 PubMed] | ||
| + | ** induced by cellobiose ([[GmuR]]) [http://www.ncbi.nlm.nih.gov/sites/entrez/18177310 PubMed] | ||
| + | ** subject to carbon catabolite repression ([[CcpA]]) [http://www.ncbi.nlm.nih.gov/sites/entrez/18177310 PubMed] | ||
* '''Regulatory mechanism:''' | * '''Regulatory mechanism:''' | ||
| + | ** [[GmuR]]: transcription repression [http://www.ncbi.nlm.nih.gov/sites/entrez/18177310 PubMed] | ||
| + | ** [[CcpA]]: transcription repression [http://www.ncbi.nlm.nih.gov/sites/entrez/18177310 PubMed] | ||
* '''Additional information:''' | * '''Additional information:''' | ||
Revision as of 19:54, 31 May 2009
- Description: glucomannan-specific phosphotransferase system, EIIC component
| Gene name | gmuC |
| Synonyms | ydhO |
| Essential | no |
| Product | glucomannan-specific phosphotransferase system, EIIC component |
| Function | glucomannan uptake and phosphorylation |
| MW, pI | 48 kDa, 9.803 |
| Gene length, protein length | 1326 bp, 442 aa |
| Immediate neighbours | gmuA, gmuD |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 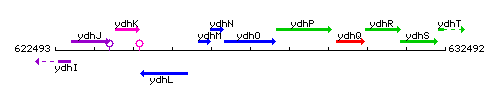
This image was kindly provided by SubtiList
| |
Contents
The gene
Basic information
- Coordinates:
Phenotypes of a mutant
Database entries
- DBTBS entry: no entry
- SubtiList entry: [1]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family: PTS permease, lactose permease (Lac) family PubMed
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Interactions:
- Localization:
Database entries
- Structure:
- Swiss prot entry:
- KEGG entry: BSU05830
- E.C. number:
Additional information
Expression and regulation
- Regulation:
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
- Reizer et al. (1999) Novel phosphotransferase system genes revealed by genome analysis - the complete complement of PTS proteins encoded within the genome of Bacillus subtilis. Microbiology 145: 3419-3429 PubMed
- Sadaie Y, Nakadate H, Fukui R, Yee LM, Asai K. (2008) Glucomannan utilization operon of Bacillus subtilis. FEMS Microbiol Lett. Feb;279(1): 103-9. PubMed
- Author1, Author2 & Author3 (year) Title Journal volume: page-page. PubMed