Difference between revisions of "Sandbox"
| Line 1: | Line 1: | ||
| − | * '''Description:''' | + | * '''Description:''' Transcriptional regulator Spx, involved in regulation of many genes. <br/><br/> |
{| align="right" border="1" cellpadding="2" | {| align="right" border="1" cellpadding="2" | ||
|- | |- | ||
|style="background:#ABCDEF;" align="center"|'''Gene name''' | |style="background:#ABCDEF;" align="center"|'''Gene name''' | ||
| − | |'' | + | |''spx'' |
|- | |- | ||
| − | |style="background:#ABCDEF;" align="center"| '''Synonyms''' || '' '' | + | |style="background:#ABCDEF;" align="center"| '''Synonyms''' || ''yjbD '' |
|- | |- | ||
| − | |style="background:#ABCDEF;" align="center"| '''Essential''' || no | + | |style="background:#ABCDEF;" align="center"| '''Essential''' || no |
|- | |- | ||
| − | |style="background:#ABCDEF;" align="center"| '''Product''' || | + | |style="background:#ABCDEF;" align="center"| '''Product''' || transcriptional regulator Spx |
|- | |- | ||
| − | |style="background:#ABCDEF;" align="center"|'''Function''' || | + | |style="background:#ABCDEF;" align="center"|'''Function''' || negative and positive regulator of many genes |
|- | |- | ||
| − | |style="background:#ABCDEF;" align="center"| '''MW, pI''' || | + | |style="background:#ABCDEF;" align="center"| '''MW, pI''' || 15,5 kDa, 7.80 |
|- | |- | ||
| − | |style="background:#ABCDEF;" align="center"| '''Gene length, protein length''' || | + | |style="background:#ABCDEF;" align="center"| '''Gene length, protein length''' || 393 bp, 131 amino acids |
|- | |- | ||
| − | |style="background:#ABCDEF;" align="center"|'''Immediate neighbours''' || ''[[ | + | |style="background:#ABCDEF;" align="center"|'''Immediate neighbours''' || ''[[yjbC]]'', ''[[yjbE]]'' |
|- | |- | ||
| − | |colspan="2" style="background:#FAF8CC;" align="center"|'''Get the DNA and protein [http://srs.ebi.ac.uk/srsbin/cgi-bin/wgetz?-e+[EMBLCDS: | + | |colspan="2" style="background:#FAF8CC;" align="center"|'''Get the DNA and protein [http://srs.ebi.ac.uk/srsbin/cgi-bin/wgetz?-e+[EMBLCDS:CAB13007]+-newId sequences] <br/> (Barbe ''et al.'', 2009)''' |
|- | |- | ||
| − | |colspan="2" | '''Genetic context''' <br/> [[Image: | + | |colspan="2" | '''Genetic context''' <br/> [[Image:spx_context.gif]] |
<div align="right"> <small>This image was kindly provided by [http://genolist.pasteur.fr/SubtiList/ SubtiList]</small></div> | <div align="right"> <small>This image was kindly provided by [http://genolist.pasteur.fr/SubtiList/ SubtiList]</small></div> | ||
|- | |- | ||
| Line 29: | Line 29: | ||
__TOC__ | __TOC__ | ||
| − | <br/><br/> | + | <br/><br/> |
| + | |||
=The gene= | =The gene= | ||
| Line 35: | Line 36: | ||
=== Basic information === | === Basic information === | ||
| − | * '''Coordinates:''' | + | * '''Coordinates:''' 1227010 - 1227402 |
===Phenotypes of a mutant === | ===Phenotypes of a mutant === | ||
| + | |||
| + | Loss of up-regulation of the methionine sulfoxide reductase (''[[mrsA]]-[[mrsB]]'') operon in response to thiol specific oxidative stress, also loss of ''[[trxA]]'' and ''[[trxB]]'' upregulation in response to thiol specific oxidative stress. | ||
=== Database entries === | === Database entries === | ||
| − | * '''DBTBS entry:''' [http://dbtbs.hgc.jp/COG/prom/ | + | * '''DBTBS entry:''' [http://dbtbs.hgc.jp/COG/prom/yjbCD.html] |
| − | * '''SubtiList entry:''' [http://genolist.pasteur.fr/SubtiList/genome.cgi?gene_detail+ | + | * '''SubtiList entry:''' [http://genolist.pasteur.fr/SubtiList/genome.cgi?gene_detail+BG13133 link] |
=== Additional information=== | === Additional information=== | ||
| − | |||
=The protein= | =The protein= | ||
| Line 52: | Line 54: | ||
=== Basic information/ Evolution === | === Basic information/ Evolution === | ||
| − | * '''Catalyzed reaction/ biological activity:''' | + | * '''Catalyzed reaction/ biological activity:''' Transcriptional regulator of many genes in response to thiol specific oxidative stress (transcription activator of'' [[trxA]]'' and ''[[trxB]]''). In addition, Spx inhibits transcription by binding to the C-terminal domain of the alpha subunit of RNAP ([[RpoA]]), disrupting complex formation between RNAP and certain transcriptional activator proteins like [[ResD]] and [[ComA]]. In response to thiol specific oxidative stress, Spx can also activate transcription, making it a general regulator that exerts both positive and negative control over transcription initiation. |
| − | * '''Protein family:''' | + | * '''Protein family:''' Arsenate Reductase (ArsC) family, Spx subfamily |
| − | * '''Paralogous protein(s):''' | + | * '''Paralogous protein(s):''' [[MgsR]] |
=== Extended information on the protein === | === Extended information on the protein === | ||
| Line 62: | Line 64: | ||
* '''Kinetic information:''' | * '''Kinetic information:''' | ||
| − | * '''Domains:''' | + | * '''Domains:''' CXXC (10-13): Acts as a disulfide switch for the redox-sensitive transcriptional regulation of genes that function in thiol homeostasis. |
| − | * '''Modification:''' | + | * '''Modification:''' Cysteine oxidation of the CXXC motif |
* '''Cofactor(s):''' | * '''Cofactor(s):''' | ||
| Line 70: | Line 72: | ||
* '''Effectors of protein activity:''' | * '''Effectors of protein activity:''' | ||
| − | * '''Interactions:''' [[ | + | * '''Interactions:''' [[RpoA]]-[[Spx]], [[YjbH]]-[[Spx]] [http://www.ncbi.nlm.nih.gov/sites/entrez/19074380 PubMed], [[Spx]]-[[RpoA]] (C-terminal domain [http://www.ncbi.nlm.nih.gov/sites/entrez/12642660 PubMed] |
| − | |||
* '''Localization:''' | * '''Localization:''' | ||
=== Database entries === | === Database entries === | ||
| − | * '''Structure:''' | + | * '''Structure:''' complex with C-terminal domain of [[RpoA]] [http://www.ncbi.nlm.nih.gov/Structure/mmdb/mmdbsrv.cgi?Dopt=s&uid=35536 NCBI] |
* '''Swiss prot entry:''' | * '''Swiss prot entry:''' | ||
| − | * '''KEGG entry:''' [http://www.genome.jp/dbget-bin/www_bget?bsu+ | + | * '''KEGG entry:''' [http://www.genome.jp/dbget-bin/www_bget?bsu+BSU11500] |
* '''E.C. number:''' | * '''E.C. number:''' | ||
=== Additional information=== | === Additional information=== | ||
| + | |||
=Expression and regulation= | =Expression and regulation= | ||
| − | * '''Operon:''' | + | * '''Operon:''' ''[[yjbC]]-[[spx]]'', ''[[spx]]'' |
| − | * '''[[Sigma factor]]:''' | + | * '''[[Sigma factor]]:''' four promoters upstream of ''[[yjbC]]'': [[SigW]], [[SigB]], [[SigX]], and unknown sigma [http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=+10913081 PubMed], |
| + | :::: promoters upstream of ''spx'': [[SigA]], [[SigW]] [http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=+10913081 PubMed], [[SigM]] [http://www.ncbi.nlm.nih.gov/sites/entrez/17434969 PubMed] | ||
| − | * '''Regulation:''' | + | * '''Regulation:''' induced by stress ([[SigB]]) [http://www.ncbi.nlm.nih.gov/pubmed/15805528 PubMed], Transcription is represed by [[PerR]] and [[YodB ]] [http://www.ncbi.nlm.nih.gov/sites/entrez/17158660 PubMed] |
| − | * '''Regulatory mechanism:''' | + | * '''Regulatory mechanism:''' transcription repression |
| − | * '''Additional information:''' | + | * '''Additional information:'''Post-translational control by [[ClpX]]-[[ClpP]]: Spx naturally contains a C-terminal sequence that resembles the [[SsrA]] tag and targets the protein for degradation. [http://www.ncbi.nlm.nih.gov/pubmed/12642660?ordinalpos=27&itool=EntrezSystem2.PEntrez.Pubmed.Pubmed_ResultsPanel.Pubmed_DefaultReportPanel.Pubmed_RVDocSum PubMed] |
| + | |||
| + | Proteolysis is enhanced by [[YjbH]]. [http://www.ncbi.nlm.nih.gov/pubmed/19074380?ordinalpos=1&itool=EntrezSystem2.PEntrez.Pubmed.Pubmed_ResultsPanel.Pubmed_DefaultReportPanel.Pubmed_RVDocSum PubMed] | ||
=Biological materials = | =Biological materials = | ||
| − | * '''Mutant:''' | + | * '''Mutant:''' ORB6781 (spc), ORB6876 (tet), available in [[Zuber]] lab |
* '''Expression vector:''' | * '''Expression vector:''' | ||
| − | + | ||
* '''lacZ fusion:''' | * '''lacZ fusion:''' | ||
* '''GFP fusion:''' | * '''GFP fusion:''' | ||
| − | * '''two-hybrid system:''' | + | * '''two-hybrid system:''' ''B. pertussis'' adenylate cyclase-based bacterial two hybrid system ([[BACTH]]), available in [[Stülke]] lab |
* '''Antibody:''' | * '''Antibody:''' | ||
| Line 114: | Line 119: | ||
=Labs working on this gene/protein= | =Labs working on this gene/protein= | ||
| − | [[ | + | [[Peter Zuber]], Oregon Health and Science University, USA |
| − | [http:// | + | [http://www.ogi.edu/people/dsp_person.cfm?person_id=411D6801-2A56-D16D-58A06B4480EDB9C7 Homepage] |
| + | |||
| + | [[Richard Brennan]], Houston, Texas, USA [http://www.mdanderson.org/departments/biochem/display.cfm?id=556ef368-6c81-4043-b74f350d41dd06cb&method=displayfull&pn=a8427ebd-d0ff-11d4-80fd00508b603a14 Homepage] | ||
=Your additional remarks= | =Your additional remarks= | ||
| Line 121: | Line 128: | ||
=References= | =References= | ||
| − | # | + | # Höper et al. (2005) Comprehensive Characterization of the Contribution of Individual [[SigB]]-Dependent General Stress Genes to Stress Resistance of ''Bacillus subtilis''. J. Bact. '''187:''' 2810-2826 [http://www.ncbi.nlm.nih.gov/pubmed/15805528 PubMed] |
| − | # | + | # Choi, S. Y., D. Reyes, M. Leelakriangsak, and P. Zuber. 2006. The global regulator Spx functions in the control of organosulfur metabolism in Bacillus subtilis. J. Bacteriol. 188:5741-5751. [http://www.ncbi.nlm.nih.gov/sites/entrez/16885442 PubMed] |
| − | # | + | # Eiamphungporn, W., and J. D. Helmann. 2008. The Bacillus subtilis sigma(M) regulon and its contribution to cell envelope stress responses. Mol. Microbiol. 67:830-848. [http://www.ncbi.nlm.nih.gov/sites/entrez/18179421 PubMed] |
| − | # | + | # Garg et al. 2009. The YjbH protein of Bacillus subtilis enhances ClpXP-catalyzed proteolysis of Spx. J. Bacteriol. 191: 1268-1277. [http://www.ncbi.nlm.nih.gov/sites/entrez/19074380 PubMed] |
| − | # | + | # Jervis et al. 2007. SigM-responsive genes of ''Bacillus subtilis'' and their promoters. J. Bacteriol. 189: 4534-4538. [http://www.ncbi.nlm.nih.gov/sites/entrez/17434969 PubMed] |
| − | # | + | # Larsson, J. T., A. Rogstam, and C. von Wachenfeldt. 2007. YjbH is a novel negative effector of the disulphide stress regulator, Spx, in Bacillus subtilis. Mol. Microbiol. 66:669-684. [http://www.ncbi.nlm.nih.gov/sites/entrez/17908206 PubMed] |
| − | # | + | # Leelakriangsak, M., K. Kobayashi, and P. Zuber. 2007. Dual negative control of spx transcription initiation from the P3 promoter by repressors PerR and YodB in Bacillus subtilis. J. Bacteriol. 189:1736-1744. [http://www.ncbi.nlm.nih.gov/sites/entrez/17158660 PubMed] |
| + | # Nakano, M. M., F. Hajarizadeh, Y. Zhu, and P. Zuber. 2001. Loss-of-function mutations in yjbD result in ClpX- and ClpP-independent competence development of Bacillus subtilis. Mol. Microbiol. 42:383-394.[http://www.ncbi.nlm.nih.gov/sites/entrez/11703662 PubMed] | ||
| + | # Nakano, S., K. N. Erwin, M. Ralle, and P. Zuber. 2005. Redox-sensitive transcriptional control by a thiol/disulphide switch in the global regulator, Spx. Mol. Microbiol. 55:498-510. [http://www.ncbi.nlm.nih.gov/sites/entrez/15659166 PubMed] | ||
| + | # Nakano, S., E. Küster-Schöck, A. D. Grossman, and P. Zuber. 2003. Spx dependent global transcriptional control is induced by thiol-specific oxidative stress in Bacillus subtilis. Proc. Natl. Acad. Sci. USA 100:13603-13608. [http://www.ncbi.nlm.nih.gov/sites/entrez/14597697 PubMed] | ||
| + | # Nakano, S., M. M. Nakano, Y. Zhang, M. Leelakriangsak, and P. Zuber. 2003. A regulatory protein that interferes with activator-stimulated transcription in bacteria. Proc. Natl. Acad. Sci. USA 100:4233-4238. [http://www.ncbi.nlm.nih.gov/sites/entrez/12642660 PubMed] | ||
| + | # Nakano, S., G. Zheng, M. M. Nakano, and P. Zuber. 2002. Multiple pathways of Spx (YjbD) proteolysis in Bacillus subtilis. J. Bacteriol. 184:3664-3670. [http://www.ncbi.nlm.nih.gov/sites/entrez/12057962 PubMed] | ||
| + | # Newberry, K. J., S. Nakano, P. Zuber, and R. G. Brennan. 2005. Crystal structure of the Bacillus subtilis anti-alpha, global transcriptional regulator, Spx, in complex with the alpha C-terminal domain of RNA polymerase. Proc. Natl. Acad. Sci. USA 102:15839-15844. [http://www.ncbi.nlm.nih.gov/sites/entrez/16249335 PubMed] | ||
| + | # Petersohn, A., J. Bernhardt, U. Gerth, D. Hoper, T. Koburger, U. Volker, and M. Hecker. 1999. Identification of sigma(B)-dependent genes in Bacillus subtilis using a promoter consensus-directed search and oligonucleotide hybridization. J. Bacteriol. 181:5718-5724. [http://www.ncbi.nlm.nih.gov/sites/entrez/10482513 PubMed] | ||
| + | # Reyes, D. Y. and P. Zuber. 2008. Activation of transcription initiation by Spx: formation of a transcription complex and identification of a cis-acting element required for transcriptional activation. Mol. Microbiol. 69:765-779. [http://www.ncbi.nlm.nih.gov/sites/entrez/18687074 PubMed] | ||
| + | # Thackray, P. D., and A. Moir. 2003. SigM, an extracytoplasmic function [[sigma factor]] of Bacillus subtilis, is activated in response to cell wall antibiotics, ethanol, heat, acid, and superoxide stress. J. Bacteriol. 185:3491-3498. [http://www.ncbi.nlm.nih.gov/sites/entrez/12775685 PubMed] | ||
| + | # Zhang, Y., S. Nakano, S. Y. Choi, and P. Zuber. 2006. Mutational analysis of the Bacillus subtilis RNA polymerase alpha C-terminal domain supports the interference model of Spx-dependent repression. J. Bacteriol. 188:4300-4311. [http://www.ncbi.nlm.nih.gov/sites/entrez/16740936 PubMed] | ||
| + | # Zhang, Y., and P. Zuber. 2007. Requirement of the zinc-binding domain of ClpX for Spx proteolysis in Bacillus subtilis and effects of disulfide stress on ClpXP activity. J. Bacteriol. 189:7669-7680. [http://www.ncbi.nlm.nih.gov/sites/entrez/17827297 PubMed] | ||
| + | # Zuber, P. 2004. Spx-RNA polymerase interaction and global transcriptional control during oxidative stress. J. Bacteriol. 186:1911-1918. [http://www.ncbi.nlm.nih.gov/sites/entrez/15028674 PubMed] | ||
Revision as of 23:53, 27 April 2009
- Description: Transcriptional regulator Spx, involved in regulation of many genes.
| Gene name | spx |
| Synonyms | yjbD |
| Essential | no |
| Product | transcriptional regulator Spx |
| Function | negative and positive regulator of many genes |
| MW, pI | 15,5 kDa, 7.80 |
| Gene length, protein length | 393 bp, 131 amino acids |
| Immediate neighbours | yjbC, yjbE |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 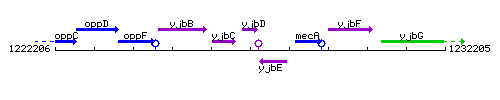
This image was kindly provided by SubtiList
| |
Contents
The gene
Basic information
- Coordinates: 1227010 - 1227402
Phenotypes of a mutant
Loss of up-regulation of the methionine sulfoxide reductase (mrsA-mrsB) operon in response to thiol specific oxidative stress, also loss of trxA and trxB upregulation in response to thiol specific oxidative stress.
Database entries
- DBTBS entry: [1]
- SubtiList entry: link
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: Transcriptional regulator of many genes in response to thiol specific oxidative stress (transcription activator of trxA and trxB). In addition, Spx inhibits transcription by binding to the C-terminal domain of the alpha subunit of RNAP (RpoA), disrupting complex formation between RNAP and certain transcriptional activator proteins like ResD and ComA. In response to thiol specific oxidative stress, Spx can also activate transcription, making it a general regulator that exerts both positive and negative control over transcription initiation.
- Protein family: Arsenate Reductase (ArsC) family, Spx subfamily
- Paralogous protein(s): MgsR
Extended information on the protein
- Kinetic information:
- Domains: CXXC (10-13): Acts as a disulfide switch for the redox-sensitive transcriptional regulation of genes that function in thiol homeostasis.
- Modification: Cysteine oxidation of the CXXC motif
- Cofactor(s):
- Effectors of protein activity:
Database entries
- Swiss prot entry:
- KEGG entry: [2]
- E.C. number:
Additional information
Expression and regulation
- Regulatory mechanism: transcription repression
- Additional information:Post-translational control by ClpX-ClpP: Spx naturally contains a C-terminal sequence that resembles the SsrA tag and targets the protein for degradation. PubMed
Proteolysis is enhanced by YjbH. PubMed
Biological materials
- Mutant: ORB6781 (spc), ORB6876 (tet), available in Zuber lab
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system: B. pertussis adenylate cyclase-based bacterial two hybrid system (BACTH), available in Stülke lab
- Antibody:
Labs working on this gene/protein
Peter Zuber, Oregon Health and Science University, USA Homepage
Richard Brennan, Houston, Texas, USA Homepage
Your additional remarks
References
- Höper et al. (2005) Comprehensive Characterization of the Contribution of Individual SigB-Dependent General Stress Genes to Stress Resistance of Bacillus subtilis. J. Bact. 187: 2810-2826 PubMed
- Choi, S. Y., D. Reyes, M. Leelakriangsak, and P. Zuber. 2006. The global regulator Spx functions in the control of organosulfur metabolism in Bacillus subtilis. J. Bacteriol. 188:5741-5751. PubMed
- Eiamphungporn, W., and J. D. Helmann. 2008. The Bacillus subtilis sigma(M) regulon and its contribution to cell envelope stress responses. Mol. Microbiol. 67:830-848. PubMed
- Garg et al. 2009. The YjbH protein of Bacillus subtilis enhances ClpXP-catalyzed proteolysis of Spx. J. Bacteriol. 191: 1268-1277. PubMed
- Jervis et al. 2007. SigM-responsive genes of Bacillus subtilis and their promoters. J. Bacteriol. 189: 4534-4538. PubMed
- Larsson, J. T., A. Rogstam, and C. von Wachenfeldt. 2007. YjbH is a novel negative effector of the disulphide stress regulator, Spx, in Bacillus subtilis. Mol. Microbiol. 66:669-684. PubMed
- Leelakriangsak, M., K. Kobayashi, and P. Zuber. 2007. Dual negative control of spx transcription initiation from the P3 promoter by repressors PerR and YodB in Bacillus subtilis. J. Bacteriol. 189:1736-1744. PubMed
- Nakano, M. M., F. Hajarizadeh, Y. Zhu, and P. Zuber. 2001. Loss-of-function mutations in yjbD result in ClpX- and ClpP-independent competence development of Bacillus subtilis. Mol. Microbiol. 42:383-394.PubMed
- Nakano, S., K. N. Erwin, M. Ralle, and P. Zuber. 2005. Redox-sensitive transcriptional control by a thiol/disulphide switch in the global regulator, Spx. Mol. Microbiol. 55:498-510. PubMed
- Nakano, S., E. Küster-Schöck, A. D. Grossman, and P. Zuber. 2003. Spx dependent global transcriptional control is induced by thiol-specific oxidative stress in Bacillus subtilis. Proc. Natl. Acad. Sci. USA 100:13603-13608. PubMed
- Nakano, S., M. M. Nakano, Y. Zhang, M. Leelakriangsak, and P. Zuber. 2003. A regulatory protein that interferes with activator-stimulated transcription in bacteria. Proc. Natl. Acad. Sci. USA 100:4233-4238. PubMed
- Nakano, S., G. Zheng, M. M. Nakano, and P. Zuber. 2002. Multiple pathways of Spx (YjbD) proteolysis in Bacillus subtilis. J. Bacteriol. 184:3664-3670. PubMed
- Newberry, K. J., S. Nakano, P. Zuber, and R. G. Brennan. 2005. Crystal structure of the Bacillus subtilis anti-alpha, global transcriptional regulator, Spx, in complex with the alpha C-terminal domain of RNA polymerase. Proc. Natl. Acad. Sci. USA 102:15839-15844. PubMed
- Petersohn, A., J. Bernhardt, U. Gerth, D. Hoper, T. Koburger, U. Volker, and M. Hecker. 1999. Identification of sigma(B)-dependent genes in Bacillus subtilis using a promoter consensus-directed search and oligonucleotide hybridization. J. Bacteriol. 181:5718-5724. PubMed
- Reyes, D. Y. and P. Zuber. 2008. Activation of transcription initiation by Spx: formation of a transcription complex and identification of a cis-acting element required for transcriptional activation. Mol. Microbiol. 69:765-779. PubMed
- Thackray, P. D., and A. Moir. 2003. SigM, an extracytoplasmic function sigma factor of Bacillus subtilis, is activated in response to cell wall antibiotics, ethanol, heat, acid, and superoxide stress. J. Bacteriol. 185:3491-3498. PubMed
- Zhang, Y., S. Nakano, S. Y. Choi, and P. Zuber. 2006. Mutational analysis of the Bacillus subtilis RNA polymerase alpha C-terminal domain supports the interference model of Spx-dependent repression. J. Bacteriol. 188:4300-4311. PubMed
- Zhang, Y., and P. Zuber. 2007. Requirement of the zinc-binding domain of ClpX for Spx proteolysis in Bacillus subtilis and effects of disulfide stress on ClpXP activity. J. Bacteriol. 189:7669-7680. PubMed
- Zuber, P. 2004. Spx-RNA polymerase interaction and global transcriptional control during oxidative stress. J. Bacteriol. 186:1911-1918. PubMed