Difference between revisions of "Fnr"
| Line 24: | Line 24: | ||
|style="background:#ABCDEF;" align="center"|'''Immediate neighbours''' || ''[[ywiC]]'', ''[[narK]]'' | |style="background:#ABCDEF;" align="center"|'''Immediate neighbours''' || ''[[ywiC]]'', ''[[narK]]'' | ||
|- | |- | ||
| − | |style="background:#FAF8CC;" align="center"|'''Sequences'''||[http://bsubcyc.org/BSUB/sequence-aa?type=GENE&object=BSU37310 Protein] [http://bsubcyc.org/BSUB/sequence?type=GENE&object=BSU37310 DNA] [http://bsubcyc.org/BSUB/seq-selector?chromosome=CHROM-1&object=BSU37310 | + | |style="background:#FAF8CC;" align="center"|'''Sequences'''||[http://bsubcyc.org/BSUB/sequence-aa?type=GENE&object=BSU37310 Protein] [http://bsubcyc.org/BSUB/sequence?type=GENE&object=BSU37310 DNA] [http://bsubcyc.org/BSUB/seq-selector?chromosome=CHROM-1&object=BSU37310 DNA_with_flanks] |
|- | |- | ||
|colspan="2" | '''Genetic context''' <br/> [[Image:fnr_context.gif]] | |colspan="2" | '''Genetic context''' <br/> [[Image:fnr_context.gif]] | ||
Revision as of 12:32, 14 May 2013
- Description: transcriptional regulator of anaerobic genes
| Gene name | fnr |
| Synonyms | |
| Essential | no |
| Product | transcription regulator (Crp family) |
| Function | regulation of anaerobiosis, fermentation and overflow metabolism |
| Gene expression levels in SubtiExpress: fnr | |
| Metabolic function and regulation of this protein in SubtiPathways: Alternative nitrogen sources | |
| MW, pI | 27 kDa, 5.989 |
| Gene length, protein length | 714 bp, 238 aa |
| Immediate neighbours | ywiC, narK |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 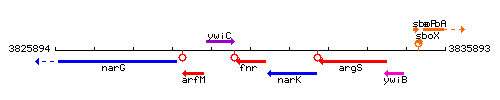
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed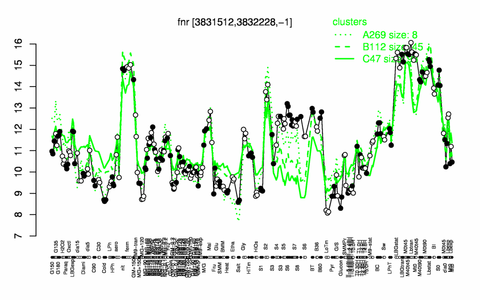
| |
Contents
Categories containing this gene/protein
regulators of electron transport, transcription factors and their control
This gene is a member of the following regulons
Fnr regulon, NsrR regulon, ResD regulon
The Fnr regulon
The gene
Basic information
- Locus tag: BSU37310
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family:
- Paralogous protein(s):
Genes/ operons controlled by Fnr
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity: contains an iron-sulfur cluster
- Localization: cytoplasm (according to Swiss-Prot)
Database entries
- Structure:
- UniProt: P46908
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Regulation:
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Reviews
The Fnr regulon
Other original publications
Ines Gruner, Claudia Frädrich, Lars H Böttger, Alfred X Trautwein, Dieter Jahn, Elisabeth Härtig
Aspartate 141 is the fourth ligand of the oxygen-sensing [4Fe-4S]2+ cluster of Bacillus subtilis transcriptional regulator Fnr.
J Biol Chem: 2011, 286(3);2017-21
[PubMed:21068385]
[WorldCat.org]
[DOI]
(I p)
Michiko M Nakano, Hao Geng, Shunji Nakano, Kazuo Kobayashi
The nitric oxide-responsive regulator NsrR controls ResDE-dependent gene expression.
J Bacteriol: 2006, 188(16);5878-87
[PubMed:16885456]
[WorldCat.org]
[DOI]
(P p)
M M Nakano, Y Zhu, M Lacelle, X Zhang, F M Hulett
Interaction of ResD with regulatory regions of anaerobically induced genes in Bacillus subtilis.
Mol Microbiol: 2000, 37(5);1198-207
[PubMed:10972836]
[WorldCat.org]
[DOI]
(P p)
M M Nakano, P Zuber, P Glaser, A Danchin, F M Hulett
Two-component regulatory proteins ResD-ResE are required for transcriptional activation of fnr upon oxygen limitation in Bacillus subtilis.
J Bacteriol: 1996, 178(13);3796-802
[PubMed:8682783]
[WorldCat.org]
[DOI]
(P p)
H Cruz Ramos, L Boursier, I Moszer, F Kunst, A Danchin, P Glaser
Anaerobic transcription activation in Bacillus subtilis: identification of distinct FNR-dependent and -independent regulatory mechanisms.
EMBO J: 1995, 14(23);5984-94
[PubMed:8846791]
[WorldCat.org]
[DOI]
(P p)