Difference between revisions of "Eno"
(→Biological materials) |
|||
| Line 152: | Line 152: | ||
* '''Mutant:''' | * '''Mutant:''' | ||
| − | ** GP594 (''eno''::''cat''), available in [[Stülke]] lab | + | ** GP594 (''eno''::''cat''), available in [[Jörg Stülke]]'s lab |
| − | ** GP599 (''eno''::''erm''), available in [[Stülke]] lab | + | ** GP599 (''eno''::''erm''), available in [[Jörg Stülke]]'s lab |
| − | ** GP698 (''eno''-''[[pgm]]''::''cat''), available in [[Stülke]] | + | ** GP698 (''eno''-''[[pgm]]''::''cat''), available in [[Jörg Stülke]]'s lab |
| − | |||
* '''Expression vector:''' | * '''Expression vector:''' | ||
| − | ** pGP1426 (expression of ''[[eno]]'' in ''B. subtilis'', in [[pBQ200]]), available in [[Stülke]] lab | + | ** pGP1426 (expression of ''[[eno]]'' in ''B. subtilis'', in [[pBQ200]]), available in [[Jörg Stülke]]'s lab |
| − | ** pGP1500 (expression of ''[[pgm]]'' and ''[[eno]]'' in ''B. subtilis'', in [[pBQ200]]), available in [[Stülke]] lab | + | ** pGP1500 (expression of ''[[pgm]]'' and ''[[eno]]'' in ''B. subtilis'', in [[pBQ200]]), available in [[Jörg Stülke]]'s lab |
| − | ** pGP563 (N-terminal His-tag, in [[pWH844]]), available in [[Stülke]] lab | + | ** pGP563 (N-terminal His-tag, in [[pWH844]]), available in [[Jörg Stülke]]'s lab |
| − | ** pGP1276 (N-terminal Strep-tag, purification from ''E. coli'', in [[pGP172]]), available in [[Stülke]] lab | + | ** pGP1276 (N-terminal Strep-tag, purification from ''E. coli'', in [[pGP172]]), available in [[Jörg Stülke]]'s lab |
| − | ** pGP93 (N-terminal Strep-tag, purification from ''B. subtilis'', for [[SPINE]], in [[pGP380]]), available in [[Stülke]] lab | + | ** pGP93 (N-terminal Strep-tag, purification from ''B. subtilis'', for [[SPINE]], in [[pGP380]]), available in [[Jörg Stülke]]'s lab |
| + | ** GP1215 (chromosomal ''eno''-''Strep'' fusion, ''spc''), purification from ''B. subtilis'', for [[SPINE]], available in [[Jörg Stülke]]'s lab | ||
* '''lacZ fusion:''' | * '''lacZ fusion:''' | ||
** see ''[[pgk]]'' | ** see ''[[pgk]]'' | ||
| − | * '''GFP fusion:''' pHT315-yfp-eno, available in [[Mijakovic]] lab | + | * '''GFP fusion:''' |
| + | ** pHT315-yfp-eno, available in [[Mijakovic]] lab | ||
| − | * '''two-hybrid system:''' ''B. pertussis'' adenylate cyclase-based bacterial two hybrid system ([[BACTH]]), available in [[Stülke]] lab | + | * '''two-hybrid system:''' ''B. pertussis'' adenylate cyclase-based bacterial two hybrid system ([[BACTH]]), available in [[Jörg Stülke]]'s lab |
| − | * '''FLAG-tag construct:''' GP1214 (spc, based on [[pGP1331]]), available in | + | * '''FLAG-tag construct:''' |
| + | ** GP1214 (spc, based on [[pGP1331]]), available in [[Jörg Stülke]]'s lab | ||
| − | * '''Antibody:''' available in [[Stülke]] lab | + | * '''Antibody:''' available in [[Jörg Stülke]]'s lab |
=Labs working on this gene/protein= | =Labs working on this gene/protein= | ||
Revision as of 13:21, 1 October 2012
- Description: enolase, glycolytic/ gluconeogenic enzyme, universally conserved protein
| Gene name | eno |
| Synonyms | |
| Essential | no |
| Product | enolase |
| Function | enzyme in glycolysis/ gluconeogenesis |
| Gene expression levels in SubtiExpress: eno | |
| Interactions involving this protein in SubtInteract: Eno | |
| Metabolic function and regulation of this protein in SubtiPathways: Central C-metabolism | |
| MW, pI | 46,4 kDa, 4.49 |
| Gene length, protein length | 1290 bp, 430 amino acids |
| Immediate neighbours | yvbK, pgm |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 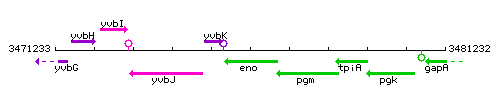
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed
| |
Contents
Categories containing this gene/protein
carbon core metabolism, membrane proteins, phosphoproteins, universally conserved proteins
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU33900
Phenotypes of a mutant
- no growth on LB, requires glucose and malate
- essential according to Kobayashi et al. on LB PubMed
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: 2-phospho-D-glycerate = phosphoenolpyruvate + H2O (according to Swiss-Prot) 2-phospho-D-glycerate = phosphoenolpyruvate + H(2)O
- Protein family: enolase family (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information: reversible Michaelis-Menten PubMed
- Domains:
- substrate binding domain (366–369)
- Cofactor(s): Mg2+
- Effectors of protein activity:
- Inhibited by EDTA PubMed
Database entries
- UniProt: P37869
- KEGG entry: [3]
- E.C. number: 4.2.1.11
Additional information
- Enolase is a moonlighting protein. PubMed
- There are indications that this enzyme is an octamer PubMed
- universally conserved protein
- extensive information on the structure and enzymatic properties of Eno can be found at Proteopedia
Expression and regulation
- Regulation:
- Additional information:
Biological materials
- Mutant:
- GP594 (eno::cat), available in Jörg Stülke's lab
- GP599 (eno::erm), available in Jörg Stülke's lab
- GP698 (eno-pgm::cat), available in Jörg Stülke's lab
- Expression vector:
- pGP1426 (expression of eno in B. subtilis, in pBQ200), available in Jörg Stülke's lab
- pGP1500 (expression of pgm and eno in B. subtilis, in pBQ200), available in Jörg Stülke's lab
- pGP563 (N-terminal His-tag, in pWH844), available in Jörg Stülke's lab
- pGP1276 (N-terminal Strep-tag, purification from E. coli, in pGP172), available in Jörg Stülke's lab
- pGP93 (N-terminal Strep-tag, purification from B. subtilis, for SPINE, in pGP380), available in Jörg Stülke's lab
- GP1215 (chromosomal eno-Strep fusion, spc), purification from B. subtilis, for SPINE, available in Jörg Stülke's lab
- lacZ fusion:
- see pgk
- GFP fusion:
- pHT315-yfp-eno, available in Mijakovic lab
- two-hybrid system: B. pertussis adenylate cyclase-based bacterial two hybrid system (BACTH), available in Jörg Stülke's lab
- FLAG-tag construct:
- GP1214 (spc, based on pGP1331), available in Jörg Stülke's lab
- Antibody: available in Jörg Stülke's lab
Labs working on this gene/protein
Jörg Stülke, University of Göttingen, Germany Homepage
Your additional remarks
References
Reviews
Subcellular localization of enolase
Additional publications: PubMed
Other original publications