Difference between revisions of "GalE"
(→Phenotypes of a mutant) |
|||
| Line 54: | Line 54: | ||
===Phenotypes of a mutant === | ===Phenotypes of a mutant === | ||
| − | + | * smooth colonies on MsGG medium, no [[biofilm formation]] in the absence of externally provided galactose {{PubMed|22893383,22113911}} | |
| − | + | * the mutant is sensitive to exogenous galactose due to the accumulation of toxic UDP-galactose, this sensitivity can be suppressed by the inactivation of ''[[galK]]'' (no formation of the toxic UDP-galactose) or of ''[[sinR]]'' (constitutive expression of the ''[[epsA]]-[[epsB]]-[[epsC]]-[[epsD]]-[[epsE]]-[[epsF]]-[[epsG]]-[[epsH]]-[[epsI]]-[[epsJ]]-[[epsK]]-[[epsL]]-[[epsM]]-[[epsN]]-[[epsO]]'' results in detoxification of UDP-galactose) {{PubMed|22893383}} | |
=== Database entries === | === Database entries === | ||
Revision as of 15:10, 16 August 2012
- Description: UDP glucose 4-epimerase
| Gene name | galE |
| Synonyms | |
| Essential | no |
| Product | UDP glucose 4-epimerase |
| Function | galactose utilization |
| Gene expression levels in SubtiExpress: galE | |
| Metabolic function and regulation of this protein in SubtiPathways: Sugar catabolism | |
| MW, pI | 36 kDa, 4.776 |
| Gene length, protein length | 1017 bp, 339 aa |
| Immediate neighbours | yxkC, yxkA |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 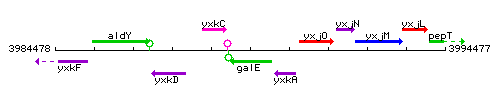
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed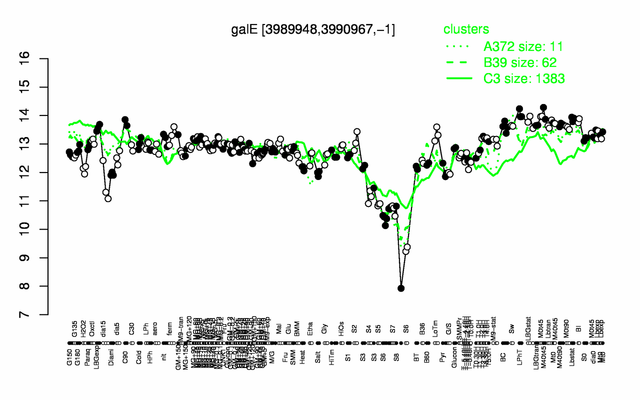
| |
Contents
Categories containing this gene/protein
utilization of specific carbon sources, biofilm formation
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU38860
Phenotypes of a mutant
- smooth colonies on MsGG medium, no biofilm formation in the absence of externally provided galactose PubMed
- the mutant is sensitive to exogenous galactose due to the accumulation of toxic UDP-galactose, this sensitivity can be suppressed by the inactivation of galK (no formation of the toxic UDP-galactose) or of sinR (constitutive expression of the epsA-epsB-epsC-epsD-epsE-epsF-epsG-epsH-epsI-epsJ-epsK-epsL-epsM-epsN-epsO results in detoxification of UDP-galactose) PubMed
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: UDP-glucose = UDP-galactose (according to Swiss-Prot)
- Protein family: sugar epimerase family (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
Database entries
- UniProt: P55180
- KEGG entry: [3]
- E.C. number: 5.1.3.2
Additional information
Expression and regulation
- Operon: galE PubMed
- Regulation:
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Additional publications: PubMed