Difference between revisions of "SwrAA/1"
(→Extended information on the protein) |
|||
| Line 1: | Line 1: | ||
| − | * '''Description:''' control of | + | * '''Description:''' control of [[DegU]] activity, enhances ''[[sigD]]'' transcription, inactive pseudogene in strain 168 <br/><br/> |
{| align="right" border="1" cellpadding="2" | {| align="right" border="1" cellpadding="2" | ||
| Line 12: | Line 12: | ||
|style="background:#ABCDEF;" align="center"| '''Product''' || swarming motility protein | |style="background:#ABCDEF;" align="center"| '''Product''' || swarming motility protein | ||
|- | |- | ||
| − | |style="background:#ABCDEF;" align="center"|'''Function''' || | + | |style="background:#ABCDEF;" align="center"|'''Function''' || control of [[DegU]] activity |
| + | |- | ||
| + | |colspan="2" style="background:#FAF8CC;" align="center"| '''Interactions involving this protein in [http://cellpublisher.gobics.de/subtinteract/startpage/start/ ''Subt''Interact]''': [http://cellpublisher.gobics.de/subtinteract/interactionList/2/SwrA SwrA] | ||
|- | |- | ||
|style="background:#ABCDEF;" align="center"| '''MW, pI''' || - , - | |style="background:#ABCDEF;" align="center"| '''MW, pI''' || - , - | ||
| Line 35: | Line 37: | ||
{{SubtiWiki category|[[biofilm formation]]}}, | {{SubtiWiki category|[[biofilm formation]]}}, | ||
{{SubtiWiki category|[[motility and chemotaxis]]}}, | {{SubtiWiki category|[[motility and chemotaxis]]}}, | ||
| + | {{SubtiWiki category|[[transcription factors and their control]]}}, | ||
{{SubtiWiki category|[[pseudogenes]]}} | {{SubtiWiki category|[[pseudogenes]]}} | ||
| Line 66: | Line 69: | ||
* '''Catalyzed reaction/ biological activity:''' | * '''Catalyzed reaction/ biological activity:''' | ||
| − | + | ** interacts with the N-terminal domain of [[DegU]] to control the activity of [[DegU]] {{PubMed|22496484}} | |
* '''Protein family:''' | * '''Protein family:''' | ||
| Line 135: | Line 138: | ||
==Original publications== | ==Original publications== | ||
| − | <pubmed>16357223,19389763,16030230,18567663,15066026,, 19389763 19749039 21278284 16091050 21602220 </pubmed> | + | <pubmed>16357223,19389763,16030230,18567663,15066026,, 19389763 19749039 21278284 16091050 21602220 22496484</pubmed> |
[[Category:Pseudogenes]] | [[Category:Pseudogenes]] | ||
Revision as of 15:48, 13 April 2012
- Description: control of DegU activity, enhances sigD transcription, inactive pseudogene in strain 168
| Gene name | swrA |
| Synonyms | yvzD, swrAA, ifm |
| Essential | no |
| Product | swarming motility protein |
| Function | control of DegU activity |
| Interactions involving this protein in SubtInteract: SwrA | |
| MW, pI | - , - |
| Gene length, protein length | 336 bp, - |
| Immediate neighbours | minJ, swrAA/2 |
| Gene sequence (+200bp) | Protein sequence |
Genetic context 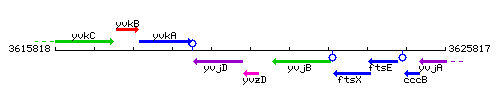
This image was kindly provided by SubtiList
| |
Contents
Categories containing this gene/protein
biofilm formation, motility and chemotaxis, transcription factors and their control, pseudogenes
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU35230
Phenotypes of a mutant
Database entries
- DBTBS entry: no entry
- SubtiList entry: [1]
Additional information
- This protein is functional in undomesticated strains of B.subtilis but not in laboratory strains, such as 168, because of a frameshift mutation. Therefore laboratory strains of B.subtilis are unable to swarm.
- The frameshift in strain 168 is caused by a single insertion of an adenine in the codon for Tyr-12 which leads to the premature truncation of the protein in residue 13. In addition, the C-terminal section of swrAA was predicted to be an ORF (yvzD) by the genome project.
- Correction of sfp, epsC, swrAA, and degQ as well as introduction of rapP from a plasmid present in NCIB3610 results in biofilm formation in B. subtilis 168 PubMed
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family:
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Localization: cytoplasm (according to Swiss-Prot)
Database entries
- Structure:
- Swiss prot entry: O32266
- KEGG entry: [2]
- E.C. number:
Additional information
Expression and regulation
- Regulation:
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Reviews
Original publications
Mitsuo Ogura, Kensuke Tsukahara
SwrA regulates assembly of Bacillus subtilis DegU via its interaction with N-terminal domain of DegU.
J Biochem: 2012, 151(6);643-55
[PubMed:22496484]
[WorldCat.org]
[DOI]
(I p)
Kassem Hamze, Sabine Autret, Krzysztof Hinc, Soumaya Laalami, Daria Julkowska, Romain Briandet, Margareth Renault, Cédric Absalon, I Barry Holland, Harald Putzer, Simone J Séror
Single-cell analysis in situ in a Bacillus subtilis swarming community identifies distinct spatially separated subpopulations differentially expressing hag (flagellin), including specialized swarmers.
Microbiology (Reading): 2011, 157(Pt 9);2456-2469
[PubMed:21602220]
[WorldCat.org]
[DOI]
(I p)
Anna L McLoon, Sarah B Guttenplan, Daniel B Kearns, Roberto Kolter, Richard Losick
Tracing the domestication of a biofilm-forming bacterium.
J Bacteriol: 2011, 193(8);2027-34
[PubMed:21278284]
[WorldCat.org]
[DOI]
(I p)
Joyce E Patrick, Daniel B Kearns
Laboratory strains of Bacillus subtilis do not exhibit swarming motility.
J Bacteriol: 2009, 191(22);7129-33
[PubMed:19749039]
[WorldCat.org]
[DOI]
(I p)
Cecilia Osera, Giuseppe Amati, Cinzia Calvio, Alessandro Galizzi
SwrAA activates poly-gamma-glutamate synthesis in addition to swarming in Bacillus subtilis.
Microbiology (Reading): 2009, 155(Pt 7);2282-2287
[PubMed:19389763]
[WorldCat.org]
[DOI]
(P p)
Cinzia Calvio, Cecilia Osera, Giuseppe Amati, Alessandro Galizzi
Autoregulation of swrAA and motility in Bacillus subtilis.
J Bacteriol: 2008, 190(16);5720-8
[PubMed:18567663]
[WorldCat.org]
[DOI]
(I p)
Daniel B Kearns, Richard Losick
Cell population heterogeneity during growth of Bacillus subtilis.
Genes Dev: 2005, 19(24);3083-94
[PubMed:16357223]
[WorldCat.org]
[DOI]
(P p)
Nicola R Stanley, Beth A Lazazzera
Defining the genetic differences between wild and domestic strains of Bacillus subtilis that affect poly-gamma-dl-glutamic acid production and biofilm formation.
Mol Microbiol: 2005, 57(4);1143-58
[PubMed:16091050]
[WorldCat.org]
[DOI]
(P p)
Cinzia Calvio, Francesco Celandroni, Emilia Ghelardi, Giuseppe Amati, Sara Salvetti, Fabrizio Ceciliani, Alessandro Galizzi, Sonia Senesi
Swarming differentiation and swimming motility in Bacillus subtilis are controlled by swrA, a newly identified dicistronic operon.
J Bacteriol: 2005, 187(15);5356-66
[PubMed:16030230]
[WorldCat.org]
[DOI]
(P p)
Daniel B Kearns, Frances Chu, Rivka Rudner, Richard Losick
Genes governing swarming in Bacillus subtilis and evidence for a phase variation mechanism controlling surface motility.
Mol Microbiol: 2004, 52(2);357-69
[PubMed:15066026]
[WorldCat.org]
[DOI]
(P p)