Pel
Revision as of 17:02, 12 April 2012 by 134.76.70.252 (talk)
- Description: pectate lyase C
| Gene name | pel |
| Synonyms | |
| Essential | no |
| Product | pectate lyase C |
| Function | degradation of polygalacturonic acid |
| MW, pI | 45 kDa, 8.421 |
| Gene length, protein length | 1260 bp, 420 aa |
| Immediate neighbours | yflT, yflS |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 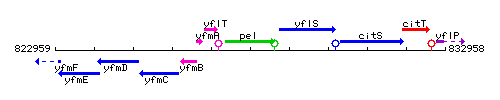
This image was kindly provided by SubtiList
| |
Contents
Categories containing this gene/protein
utilization of specific carbon sources
This gene is a member of the following regulons
CcpA regulon, ComA regulon, TnrA regulon
The gene
Basic information
- Locus tag: BSU07560
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: Eliminative cleavage of (1->4)-alpha-D-galacturonan to give oligosaccharides with 4-deoxy-alpha-D-galact-4-enuronosyl groups at their non-reducing ends (according to Swiss-Prot)
- Protein family: polysaccharide lyase 1 family (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Localization:
- extracellular (signal peptide), major constituent of the secretome PubMed
Database entries
- UniProt: P39116
- KEGG entry: [3]
- E.C. number: 4.2.2.2
Additional information
Expression and regulation
- Operon: pel PubMed
- Sigma factor:
- Regulation:
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Reviews
Massimiliano Marvasi, Pieter T Visscher, Lilliam Casillas Martinez
Exopolymeric substances (EPS) from Bacillus subtilis: polymers and genes encoding their synthesis.
FEMS Microbiol Lett: 2010, 313(1);1-9
[PubMed:20735481]
[WorldCat.org]
[DOI]
(I p)
Original publications