SpoIIAB
- Description: anti-SigF/ protein serine kinase
| Gene name | spoIIAB |
| Synonyms | |
| Essential | no |
| Product | anti-SigF/ protein serine kinase |
| Function | control of SigF activity septation; phosphorylation and inactivation of SpoIIAA |
| Gene expression levels in SubtiExpress: spoIIAB | |
| Interactions involving this protein in SubtInteract: SpoIIAB | |
| Metabolic function and regulation of this protein in SubtiPathways: spoIIAB | |
| MW, pI | 16 kDa, 4.383 |
| Gene length, protein length | 438 bp, 146 aa |
| Immediate neighbours | sigF, spoIIAA |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 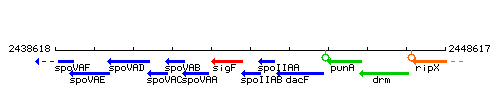
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed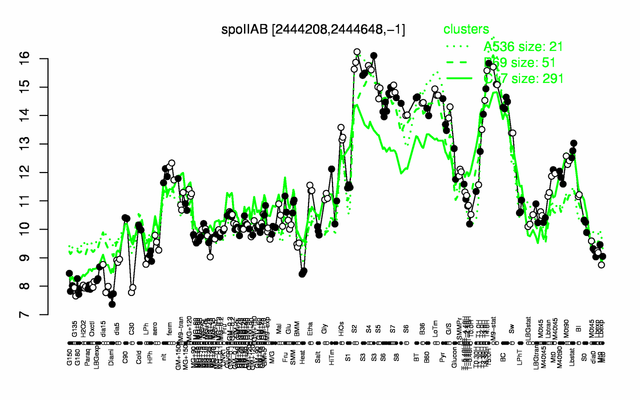
| |
Contents
Categories containing this gene/protein
protein modification, sigma factors and their control, sporulation proteins
This gene is a member of the following regulons
AbrB regulon, SigF regulon, SigG regulon, SigH regulon, SinR regulon, Spo0A regulon
The gene
Basic information
- Locus tag: BSU23460
Phenotypes of a mutant
Database entries
- BsubCyc: BSU23460
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: ATP + a protein = ADP + a phosphoprotein (according to Swiss-Prot)
- Protein family: anti-sigma-factor family (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
Database entries
- BsubCyc: BSU23460
- Structure: 1L0O (complex with SigF, Geobacillus stearothermophilus), 1TIL (complex with SpoIIAA, Geobacillus stearothermophilus)
- UniProt: P10728
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Regulation:
- Additional information:
- number of protein molecules per cell (minimal medium with glucose and ammonium): 140 PubMed
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
- Charles Moran, Emory University, NC, USA homepage
Your additional remarks
References
Modeling of SigF activation
Georgios Fengos, Dagmar Iber
Prediction stability in a data-based, mechanistic model of σF regulation during sporulation in Bacillus subtilis.
Sci Rep: 2013, 3;2755
[PubMed:24067622]
[WorldCat.org]
[DOI]
(I e)
Dagmar Iber
Inferring Biological Mechanisms by Data-Based Mathematical Modelling: Compartment-Specific Gene Activation during Sporulation in Bacillus subtilis as a Test Case.
Adv Bioinformatics: 2011, 2011;124062
[PubMed:22312331]
[WorldCat.org]
[DOI]
(I p)
Jin Hwan Do, Masao Nagasaki, Satoru Miyano
The systems approach to the prespore-specific activation of sigma factor SigF in Bacillus subtilis.
Biosystems: 2010, 100(3);178-84
[PubMed:20298743]
[WorldCat.org]
[DOI]
(I p)
Original Publications
Onuma Chumsakul, Hiroki Takahashi, Taku Oshima, Takahiro Hishimoto, Shigehiko Kanaya, Naotake Ogasawara, Shu Ishikawa
Genome-wide binding profiles of the Bacillus subtilis transition state regulator AbrB and its homolog Abh reveals their interactive role in transcriptional regulation.
Nucleic Acids Res: 2011, 39(2);414-28
[PubMed:20817675]
[WorldCat.org]
[DOI]
(I p)
Oleg A Igoshin, Chester W Price, Michael A Savageau
Signalling network with a bistable hysteretic switch controls developmental activation of the sigma transcription factor in Bacillus subtilis.
Mol Microbiol: 2006, 61(1);165-84
[PubMed:16824103]
[WorldCat.org]
[DOI]
(P p)
Masaya Fujita, José Eduardo González-Pastor, Richard Losick
High- and low-threshold genes in the Spo0A regulon of Bacillus subtilis.
J Bacteriol: 2005, 187(4);1357-68
[PubMed:15687200]
[WorldCat.org]
[DOI]
(P p)
Joanna Clarkson, Iain D Campbell, Michael D Yudkin
Efficient regulation of sigmaF, the first sporulation-specific sigma factor in B.subtilis.
J Mol Biol: 2004, 342(4);1187-95
[PubMed:15351644]
[WorldCat.org]
[DOI]
(P p)
Karen Carniol, Tae-Jong Kim, Chester W Price, Richard Losick
Insulation of the sigmaF regulatory system in Bacillus subtilis.
J Bacteriol: 2004, 186(13);4390-4
[PubMed:15205443]
[WorldCat.org]
[DOI]
(P p)
Joanna Clarkson, Iain D Campbell, Michael D Yudkin
Physical evidence for the induced release of the Bacillus subtilis transcription factor, sigma(F), from its inhibitory complex.
J Mol Biol: 2004, 340(2);203-9
[PubMed:15201047]
[WorldCat.org]
[DOI]
(P p)
Mónica Serrano, Alexandre Neves, Cláudio M Soares, Charles P Moran, Adriano O Henriques
Role of the anti-sigma factor SpoIIAB in regulation of sigmaG during Bacillus subtilis sporulation.
J Bacteriol: 2004, 186(12);4000-13
[PubMed:15175314]
[WorldCat.org]
[DOI]
(P p)
Joanna Clarkson, Jwu-Ching Shu, David A Harris, Iain D Campbell, Michael D Yudkin
Fluorescence and kinetic analysis of the SpoIIAB phosphorylation reaction, a key regulator of sporulation in Bacillus subtilis.
Biochemistry: 2004, 43(11);3120-8
[PubMed:15023063]
[WorldCat.org]
[DOI]
(P p)
Karen Carniol, Patrick Eichenberger, Richard Losick
A threshold mechanism governing activation of the developmental regulatory protein sigma F in Bacillus subtilis.
J Biol Chem: 2004, 279(15);14860-70
[PubMed:14744853]
[WorldCat.org]
[DOI]
(P p)
Qi Pan, Richard Losick
Unique degradation signal for ClpCP in Bacillus subtilis.
J Bacteriol: 2003, 185(17);5275-8
[PubMed:12923101]
[WorldCat.org]
[DOI]
(P p)
Louise Evans, Joanna Clarkson, Michael D Yudkin, Jeff Errington, Andrea Feucht
Analysis of the interaction between the transcription factor sigmaG and the anti-sigma factor SpoIIAB of Bacillus subtilis.
J Bacteriol: 2003, 185(15);4615-9
[PubMed:12867473]
[WorldCat.org]
[DOI]
(P p)
Margaret S Ho, Karen Carniol, Richard Losick
Evidence in support of a docking model for the release of the transcription factor sigma F from the antisigma factor SpoIIAB in Bacillus subtilis.
J Biol Chem: 2003, 278(23);20898-905
[PubMed:12676949]
[WorldCat.org]
[DOI]
(P p)
J Clarkson, I D Campbell, M D Yudkin
NMR studies of the interactions of SpoIIAA with its partner proteins that regulate sporulation in Bacillus subtilis.
J Mol Biol: 2001, 314(3);359-64
[PubMed:11846550]
[WorldCat.org]
[DOI]
(P p)
J Dworkin, R Losick
Differential gene expression governed by chromosomal spatial asymmetry.
Cell: 2001, 107(3);339-46
[PubMed:11701124]
[WorldCat.org]
[DOI]
(P p)
Q Pan, D A Garsin, R Losick
Self-reinforcing activation of a cell-specific transcription factor by proteolysis of an anti-sigma factor in B. subtilis.
Mol Cell: 2001, 8(4);873-83
[PubMed:11684022]
[WorldCat.org]
[DOI]
(P p)
E A Campbell, S A Darst
The anti-sigma factor SpoIIAB forms a 2:1 complex with sigma(F), contacting multiple conserved regions of the sigma factor.
J Mol Biol: 2000, 300(1);17-28
[PubMed:10864495]
[WorldCat.org]
[DOI]
(P p)
A Feucht, R A Daniel, J Errington
Characterization of a morphological checkpoint coupling cell-specific transcription to septation in Bacillus subtilis.
Mol Microbiol: 1999, 33(5);1015-26
[PubMed:10476035]
[WorldCat.org]
[DOI]
(P p)
D A Garsin, L Duncan, D M Paskowitz, R Losick
The kinase activity of the antisigma factor SpoIIAB is required for activation as well as inhibition of transcription factor sigmaF during sporulation in Bacillus subtilis.
J Mol Biol: 1998, 284(3);569-78
[PubMed:9826499]
[WorldCat.org]
[DOI]
(P p)
D A Garsin, D M Paskowitz, L Duncan, R Losick
Evidence for common sites of contact between the antisigma factor SpoIIAB and its partners SpoIIAA and the developmental transcription factor sigmaF in Bacillus subtilis.
J Mol Biol: 1998, 284(3);557-68
[PubMed:9826498]
[WorldCat.org]
[DOI]
(P p)
M Lord, T Magnin, M D Yudkin
Protein conformational change and nucleotide binding involved in regulation of sigmaF in Bacillus subtilis.
J Bacteriol: 1996, 178(23);6730-5
[PubMed:8955289]
[WorldCat.org]
[DOI]
(P p)
P J Lewis, T Magnin, J Errington
Compartmentalized distribution of the proteins controlling the prespore-specific transcription factor sigmaF of Bacillus subtilis.
Genes Cells: 1996, 1(10);881-94
[PubMed:9077448]
[WorldCat.org]
[DOI]
(P p)
A L Decatur, R Losick
Three sites of contact between the Bacillus subtilis transcription factor sigmaF and its antisigma factor SpoIIAB.
Genes Dev: 1996, 10(18);2348-58
[PubMed:8824593]
[WorldCat.org]
[DOI]
(P p)
E M Kellner, A Decatur, C P Moran
Two-stage regulation of an anti-sigma factor determines developmental fate during bacterial endospore formation.
Mol Microbiol: 1996, 21(5);913-24
[PubMed:8885263]
[WorldCat.org]
[DOI]
(P p)
S Alper, A Dufour, D A Garsin, L Duncan, R Losick
Role of adenosine nucleotides in the regulation of a stress-response transcription factor in Bacillus subtilis.
J Mol Biol: 1996, 260(2);165-77
[PubMed:8764398]
[WorldCat.org]
[DOI]
(P p)
L Duncan, S Alper, R Losick
SpoIIAA governs the release of the cell-type specific transcription factor sigma F from its anti-sigma factor SpoIIAB.
J Mol Biol: 1996, 260(2);147-64
[PubMed:8764397]
[WorldCat.org]
[DOI]
(P p)
A Feucht, T Magnin, M D Yudkin, J Errington
Bifunctional protein required for asymmetric cell division and cell-specific transcription in Bacillus subtilis.
Genes Dev: 1996, 10(7);794-803
[PubMed:8846916]
[WorldCat.org]
[DOI]
(P p)
T Magnin, M Lord, J Errington, M D Yudkin
Establishing differential gene expression in sporulating Bacillus subtilis: phosphorylation of SpoIIAA (anti-anti-sigmaF) alters its conformation and prevents formation of a SpoIIAA/SpoIIAB/ADP complex.
Mol Microbiol: 1996, 19(4);901-7
[PubMed:8820658]
[WorldCat.org]
[DOI]
(P p)
L Duncan, S Alper, F Arigoni, R Losick, P Stragier
Activation of cell-specific transcription by a serine phosphatase at the site of asymmetric division.
Science: 1995, 270(5236);641-4
[PubMed:7570023]
[WorldCat.org]
[DOI]
(P p)
S Alper, L Duncan, R Losick
An adenosine nucleotide switch controlling the activity of a cell type-specific transcription factor in B. subtilis.
Cell: 1994, 77(2);195-205
[PubMed:8168129]
[WorldCat.org]
[DOI]
(P p)
D Foulger, J Errington
Effects of new mutations in the spoIIAB gene of Bacillus subtilis on the regulation of sigma F and sigma G activities.
J Gen Microbiol: 1993, 139(12);3197-203
[PubMed:8126438]
[WorldCat.org]
[DOI]
(P p)
K T Min, C M Hilditch, B Diederich, J Errington, M D Yudkin
Sigma F, the first compartment-specific transcription factor of B. subtilis, is regulated by an anti-sigma factor that is also a protein kinase.
Cell: 1993, 74(4);735-42
[PubMed:8358793]
[WorldCat.org]
[DOI]
(P p)
P A Kirchman, H DeGrazia, E M Kellner, C P Moran
Forespore-specific disappearance of the sigma-factor antagonist spoIIAB: implications for its role in determination of cell fate in Bacillus subtilis.
Mol Microbiol: 1993, 8(4);663-71
[PubMed:8332059]
[WorldCat.org]
[DOI]
(P p)
L Duncan, R Losick
SpoIIAB is an anti-sigma factor that binds to and inhibits transcription by regulatory protein sigma F from Bacillus subtilis.
Proc Natl Acad Sci U S A: 1993, 90(6);2325-9
[PubMed:8460142]
[WorldCat.org]
[DOI]
(P p)
K York, T J Kenney, S Satola, C P Moran, H Poth, P Youngman
Spo0A controls the sigma A-dependent activation of Bacillus subtilis sporulation-specific transcription unit spoIIE.
J Bacteriol: 1992, 174(8);2648-58
[PubMed:1556084]
[WorldCat.org]
[DOI]
(P p)
R Coppolecchia, H DeGrazia, C P Moran
Deletion of spoIIAB blocks endospore formation in Bacillus subtilis at an early stage.
J Bacteriol: 1991, 173(21);6678-85
[PubMed:1938874]
[WorldCat.org]
[DOI]
(P p)
P Margolis, A Driks, R Losick
Establishment of cell type by compartmentalized activation of a transcription factor.
Science: 1991, 254(5031);562-5
[PubMed:1948031]
[WorldCat.org]
[DOI]
(P p)
R Schmidt, P Margolis, L Duncan, R Coppolecchia, C P Moran, R Losick
Control of developmental transcription factor sigma F by sporulation regulatory proteins SpoIIAA and SpoIIAB in Bacillus subtilis.
Proc Natl Acad Sci U S A: 1990, 87(23);9221-5
[PubMed:2123551]
[WorldCat.org]
[DOI]
(P p)
J Errington, J Mandelstam
Use of a lacZ gene fusion to determine the dependence pattern of sporulation operon spoIIA in spo mutants of Bacillus subtilis.
J Gen Microbiol: 1986, 132(11);2967-76
[PubMed:3114419]
[WorldCat.org]
[DOI]
(P p)