Difference between revisions of "PdxS"
(→References) |
|||
| Line 120: | Line 120: | ||
=References= | =References= | ||
| − | + | <pubmed> 17726680, 16493705, 14651647, 16030023, 14762015, 19152323 </pubmed> | |
| − | |||
| − | |||
| − | |||
| − | |||
Revision as of 19:46, 22 May 2009
- Description: pyridoxal-5'-phosphate synthase (synthase domain)
| Gene name | pdxS |
| Synonyms | yaaD |
| Essential | no |
| Product | pyridoxal-5'-phosphate synthase (synthase domain) |
| Function | pyridoxal-5'-phosphate biosynthesis |
| MW, pI | 31 kDa, 5.085 |
| Gene length, protein length | 882 bp, 294 aa |
| Immediate neighbours | dacA, pdxT |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 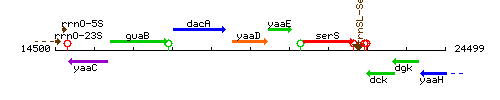
This image was kindly provided by SubtiList
| |
Contents
The gene
Basic information
- Coordinates:
Phenotypes of a mutant
auxotrophic for pyridoxal 5'-phosphate PubMed
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family: pdxS/SNZ family (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Cofactor(s):
- Effectors of protein activity:
- Localization:
Database entries
- Swiss prot entry: P37527
- KEGG entry: BSU00110
- E.C. number:
Additional information
Expression and regulation
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Silvia Wallner, Martina Neuwirth, Karlheinz Flicker, Ivo Tews, Peter Macheroux
Dissection of contributions from invariant amino acids to complex formation and catalysis in the heteromeric pyridoxal 5-phosphate synthase complex from Bacillus subtilis.
Biochemistry: 2009, 48(9);1928-35
[PubMed:19152323]
[WorldCat.org]
[DOI]
(I p)
Christine Eymann, Dörte Becher, Jörg Bernhardt, Katrin Gronau, Anja Klutzny, Michael Hecker
Dynamics of protein phosphorylation on Ser/Thr/Tyr in Bacillus subtilis.
Proteomics: 2007, 7(19);3509-26
[PubMed:17726680]
[WorldCat.org]
[DOI]
(P p)
Alain Lévine, Françoise Vannier, Cédric Absalon, Lauriane Kuhn, Peter Jackson, Elaine Scrivener, Valérie Labas, Joëlle Vinh, Patrick Courtney, Jérôme Garin, Simone J Séror
Analysis of the dynamic Bacillus subtilis Ser/Thr/Tyr phosphoproteome implicated in a wide variety of cellular processes.
Proteomics: 2006, 6(7);2157-73
[PubMed:16493705]
[WorldCat.org]
[DOI]
(P p)
Thomas Raschle, Nikolaus Amrhein, Teresa B Fitzpatrick
On the two components of pyridoxal 5'-phosphate synthase from Bacillus subtilis.
J Biol Chem: 2005, 280(37);32291-300
[PubMed:16030023]
[WorldCat.org]
[DOI]
(P p)
Boris R Belitsky
Physical and enzymological interaction of Bacillus subtilis proteins required for de novo pyridoxal 5'-phosphate biosynthesis.
J Bacteriol: 2004, 186(4);1191-6
[PubMed:14762015]
[WorldCat.org]
[DOI]
(P p)
Virginie Molle, Masaya Fujita, Shane T Jensen, Patrick Eichenberger, José E González-Pastor, Jun S Liu, Richard Losick
The Spo0A regulon of Bacillus subtilis.
Mol Microbiol: 2003, 50(5);1683-701
[PubMed:14651647]
[WorldCat.org]
[DOI]
(P p)