Difference between revisions of "PrkC"
| Line 58: | Line 58: | ||
=== Database entries === | === Database entries === | ||
| + | * '''BsubCyc:''' [http://bsubcyc.org/BSUB/NEW-IMAGE?type=NIL&object=BSU15770&redirect=T BSU15770] | ||
* '''DBTBS entry:''' no entry | * '''DBTBS entry:''' no entry | ||
| Line 96: | Line 97: | ||
=== Database entries === | === Database entries === | ||
| + | * '''BsubCyc:''' [http://bsubcyc.org/BSUB/NEW-IMAGE?type=NIL&object=BSU15770&redirect=T BSU15770] | ||
* '''Structure:''' [http://www.rcsb.org/pdb/search/structidSearch.do?structureId=3PY9 3PY3] (entire extra-cellular region of PrkC from ''Staphylococcus aureus'') {{PubMed|21208192}} | * '''Structure:''' [http://www.rcsb.org/pdb/search/structidSearch.do?structureId=3PY9 3PY3] (entire extra-cellular region of PrkC from ''Staphylococcus aureus'') {{PubMed|21208192}} | ||
Revision as of 14:41, 2 April 2014
- Description: protein kinase C, induce germination of spores in response to DAP-type, and not to Lys-type cell wall muropeptides
| Gene name | prkC |
| Synonyms | yloP |
| Essential | no |
| Product | protein kinase |
| Function | germination in response to muropeptides |
| Gene expression levels in SubtiExpress: prkC | |
| Metabolic function and regulation of this protein in SubtiPathways: prkC | |
| MW, pI | 71 kDa, 4.833 |
| Gene length, protein length | 1944 bp, 648 aa |
| Immediate neighbours | prpC, cpgA |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 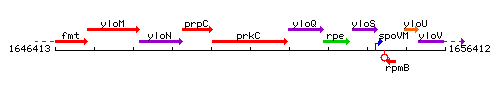
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed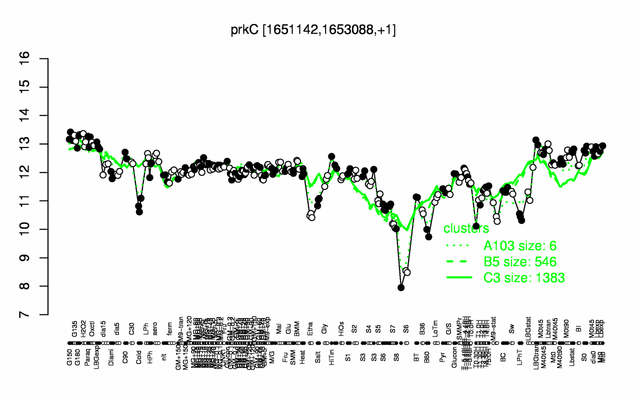
| |
Contents
Categories containing this gene/protein
protein modification, germination, membrane proteins, phosphoproteins
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU15770
Phenotypes of a mutant
- unable to germinate in response to muropeptides PubMed
Database entries
- BsubCyc: BSU15770
- DBTBS entry: no entry
- SubtiList entry: [1]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: ATP + a protein = ADP + a phosphoprotein (according to Swiss-Prot)
- Protein family: protein kinase domain (according to Swiss-Prot)
- Paralogous protein(s):
Proteins phosphorylated by PrkC
CpgA, EF-Tu, YezB PubMed, EF-G PubMed, YwjH, GlnA, Icd, AlsD, HPr PubMed, YkwC PubMed
Extended information on the protein
- Kinetic information:
- Modification: phosphorylation on Thr-290 PubMed, autophosphorylation on multiple threonine residues PubMed
- Effectors of protein activity: activated by muropeptides PubMed
- Localization: inner spore membrane PubMed, membrane PubMed
Database entries
- BsubCyc: BSU15770
- UniProt: O34507
- KEGG entry: [2]
- E.C. number: 2.7.11.1
Additional information
Expression and regulation
- Regulation:
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant: GP576 (spc), OMG302 (aphA3), available in Stülke lab
- Expression vector:
- for expression/ purification from B. subtilis with N-terminal Strep-tag, for SPINE, in pGP380: pGP832, available in Stülke lab
- for expression/ purification of the kinase domain from B. subtilis with N-terminal Strep-tag, for SPINE, in pGP380: pGP849, available in Stülke lab
- for expression, purification in E. coli with N-terminal His-tag, in pWH844: pGP1001, available in Stülke lab
- for expression, purification in E. coli with N-terminal Strep-tag, in pGP172: pGP825, available in Stülke lab
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Reviews
Phosphorylation of PrkC
Targets of PrkC-dependent phosphorylation
Phsiological role of PrkC
Expression of PrkC
Structure/ biochemistry of PrkC