Difference between revisions of "Tig"
| Line 81: | Line 81: | ||
* '''Domains:''' | * '''Domains:''' | ||
| − | * '''Modification:''' phosphorylated on ser/ thr/ tyr [http://www.ncbi.nlm.nih.gov/pubmed/16493705 PubMed] | + | * '''Modification:''' |
| + | ** phosphorylated on Arg-90 {{PubMed|22517742}} | ||
| + | ** phosphorylated on ser/ thr/ tyr [http://www.ncbi.nlm.nih.gov/pubmed/16493705 PubMed] | ||
* '''Cofactor(s):''' | * '''Cofactor(s):''' | ||
| Line 143: | Line 145: | ||
<pubmed>16231086 15763705 15837180 19647435 </pubmed> | <pubmed>16231086 15763705 15837180 19647435 </pubmed> | ||
==Original publications== | ==Original publications== | ||
| − | <pubmed>12224648,9748346,9063446,18497744 ,16493705, 8969504 20525796 </pubmed> | + | <pubmed>12224648,9748346,9063446,18497744 ,16493705, 8969504 20525796 22517742</pubmed> |
[[Category:Protein-coding genes]] | [[Category:Protein-coding genes]] | ||
Revision as of 18:43, 21 April 2012
- Description: trigger factor (prolyl isomerase)
| Gene name | tig |
| Synonyms | yzzH |
| Essential | no |
| Product | trigger factor (prolyl isomerase) |
| Function | protein folding |
| MW, pI | 47 kDa, 4.224 |
| Gene length, protein length | 1272 bp, 424 aa |
| Immediate neighbours | clpX, ysoA |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 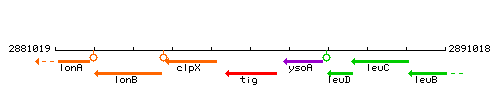
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed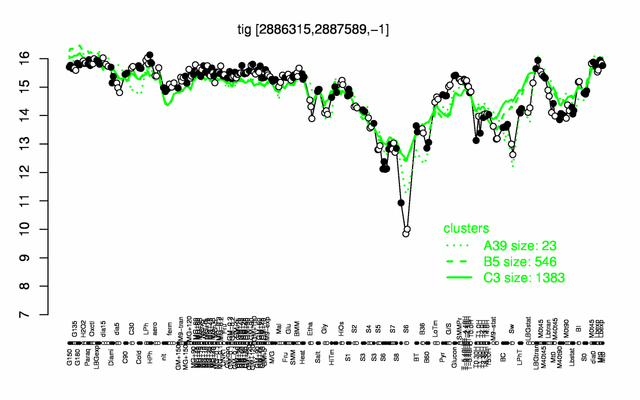
| |
Contents
Categories containing this gene/protein
chaperones/ protein folding, phosphoproteins
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU28230
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family: Tig subfamily (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Cofactor(s):
- Effectors of protein activity:
Database entries
- UniProt: P80698
- KEGG entry: [3]
- E.C. number:
Additional information
- subject to Clp-dependent proteolysis upon glucose starvation PubMed
Expression and regulation
- Operon: tig PubMed
- Sigma factor:
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Reviews
Original publications