Difference between revisions of "DivIVA"
| Line 26: | Line 26: | ||
|colspan="2" | '''Genetic context''' <br/> [[Image:divIVA_context.gif]] | |colspan="2" | '''Genetic context''' <br/> [[Image:divIVA_context.gif]] | ||
<div align="right"> <small>This image was kindly provided by [http://genolist.pasteur.fr/SubtiList/ SubtiList]</small></div> | <div align="right"> <small>This image was kindly provided by [http://genolist.pasteur.fr/SubtiList/ SubtiList]</small></div> | ||
| + | |- | ||
| + | |colspan="2" |'''[http://genome.jouy.inra.fr/cgi-bin/seb/viewdetail.py?id=divIVA_1612521_1613015_1 Expression at a glance]'''   {{PubMed|22383849}}<br/>[[Image:divIVA_expression.png|500px]] | ||
|- | |- | ||
|} | |} | ||
__TOC__ | __TOC__ | ||
| + | <br/><br/><br/><br/> | ||
| + | <br/><br/><br/><br/> | ||
| + | <br/><br/><br/><br/> | ||
| + | <br/><br/><br/><br/> | ||
| + | <br/><br/><br/><br/> | ||
| + | |||
<br/><br/> | <br/><br/> | ||
Revision as of 10:09, 19 April 2012
- Description: cell-division initiation protein (septum placement), part of the Min system (with MinC, MinD, MinJ), Noc and the Min system ensure the efficient utilization of the division site at midcell in by ensuring Z ring placement
| Gene name | divIVA |
| Synonyms | ylmJ |
| Essential | no |
| Product | cell-division initiation protein |
| Function | septum placement |
| Interactions involving this protein in SubtInteract: DivIVA | |
| MW, pI | 19 kDa, 4.846 |
| Gene length, protein length | 492 bp, 164 aa |
| Immediate neighbours | ylmH, ileS |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 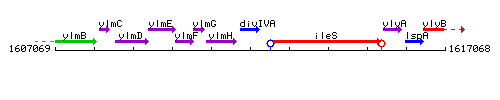
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed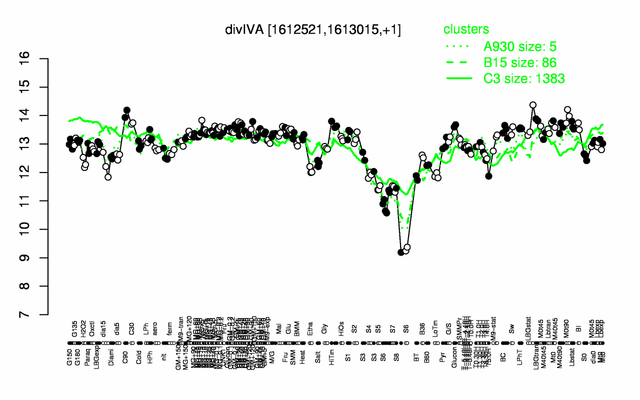
| |
Contents
Categories containing this gene/protein
cell division, membrane proteins
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU15420
Phenotypes of a mutant
Deletion of divIVA leads to filamentation and polar divisions that in turn cause a minicell phenotype. PubMed A divIVA mutant has a severe sporulation defect. PubMed
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
Filamentation is suppressed by mutations in minCD PubMed.
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: DivIVA is required for polar localisation of MinCD via MinJ. PubMed It also recruits RacA to the distal pole of the prespore PubMed.
- Protein family: gpsB family (according to Swiss-Prot)
- Paralogous protein(s): GpsB
Extended information on the protein
- Kinetic information:
- Domains: The first 60 amino acids constitute a lipid binding domain. PubMed
Multimerisation involves two coiled-coil motifs, one in the lipid binding domain, and the other one being present in the helical C-terminal domain PubMed.
- Modification: The Mycobacterium DivIVA homologue Wag31 is phosphorylated at T73 PubMed.
The DivIVA from Streptococcus pneumoniae is phosphorylated at a threonine side chain by the Ser/Thr protein kinase Sktp1. PubMed
- Cofactor(s): not known
- Effectors of protein activity: not known
Database entries
- UniProt: P71021
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Operon: divIVA PubMed
- Sigma factor:
- Additional information:
Biological materials
- Mutant: divIVA::tet available from the Hamoen Lab
- Expression vector:
- lacZ fusion:
- GFP fusion: divIVA-gfp fusions available from the Hamoen Lab
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Leendert Hamoen, Centre for Bacterial Cell Biology, Newcastle upon Tyne, United Kingdom [4]
Imrich Barak, Slovak Academy of Science, Bratislava, Slovakia homepage
Your additional remarks
References
Reviews
Original Publications