Difference between revisions of "DivIVA"
(→Extended information on the protein) |
|||
| Line 84: | Line 84: | ||
* '''Effectors of protein activity:''' not known | * '''Effectors of protein activity:''' not known | ||
| − | * '''Interactions:''' | + | * '''[[SubtInteract|Interactions]]:''' |
** ([[MinC]]-[[MinD]])-[[MinJ]]-[[DivIVA]] {{PubMed|20352045}} | ** ([[MinC]]-[[MinD]])-[[MinJ]]-[[DivIVA]] {{PubMed|20352045}} | ||
** [[DivIVA]]-[[RacA]], [[DivIVA]]-[[DivIVA]] | ** [[DivIVA]]-[[RacA]], [[DivIVA]]-[[DivIVA]] | ||
| Line 90: | Line 90: | ||
| − | * '''Localization:''' DivIVA forms a ring underneath the invaginating membrane at the site of cell division and is enriched at both cell poles [http://www.ncbi.nlm.nih.gov/sites/entrez/9219999 PubMed]. | + | * '''[[Localization]]:''' |
| + | ** DivIVA forms a ring underneath the invaginating membrane at the site of cell division and is enriched at both cell poles [http://www.ncbi.nlm.nih.gov/sites/entrez/9219999 PubMed]. | ||
=== Database entries === | === Database entries === | ||
Revision as of 13:32, 10 August 2011
- Description: cell-division initiation protein (septum placement)
| Gene name | divIVA |
| Synonyms | ylmJ |
| Essential | no |
| Product | cell-division initiation protein |
| Function | septum placement |
| Interactions involving this protein in SubtInteract: DivIVA | |
| MW, pI | 19 kDa, 4.846 |
| Gene length, protein length | 492 bp, 164 aa |
| Immediate neighbours | ylmH, ileS |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 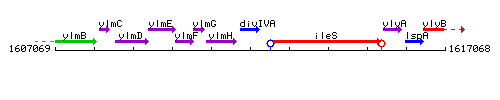
This image was kindly provided by SubtiList
| |
Contents
Categories containing this gene/protein
cell division, membrane proteins
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU15420
Phenotypes of a mutant
Deletion of divIVA leads to filamentation and polar divisions that in turn cause a minicell phenotype. PubMed A divIVA mutant has a severe sporulation defect. PubMed
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
Filamentation is suppressed by mutations in minCD PubMed.
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: DivIVA is required for polar localisation of MinCD via MinJ. PubMed It also recruits RacA to the distal pole of the prespore PubMed.
- Protein family: gpsB family (according to Swiss-Prot)
- Paralogous protein(s): GpsB
Extended information on the protein
- Kinetic information:
- Domains: The first 60 amino acids constitute a lipid binding domain. PubMed
Multimerisation involves two coiled-coil motifs, one in the lipid binding domain, and the other one being present in the helical C-terminal domain PubMed.
- Modification: The Mycobacterium DivIVA homologue Wag31 is phosphorylated at T73 PubMed.
The DivIVA from Streptococcus pneumoniae is phosphorylated at a threonine side chain by the Ser/Thr protein kinase Sktp1. PubMed
- Cofactor(s): not known
- Effectors of protein activity: not known
- Localization:
- DivIVA forms a ring underneath the invaginating membrane at the site of cell division and is enriched at both cell poles PubMed.
Database entries
- UniProt: P71021
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Operon: divIVA PubMed
- Additional information:
Biological materials
- Mutant: divIVA::tet available from the Hamoen Lab
- Expression vector:
- lacZ fusion:
- GFP fusion: divIVA-gfp fusions available from the Hamoen Lab
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Leendert Hamoen, Centre for Bacterial Cell Biology, Newcastle upon Tyne, United Kingdom [4]
Imrich Barak, Slovak Academy of Science, Bratislava, Slovakia homepage
Your additional remarks
References
Reviews
Marc Bramkamp, Suey van Baarle
Division site selection in rod-shaped bacteria.
Curr Opin Microbiol: 2009, 12(6);683-8
[PubMed:19884039]
[WorldCat.org]
[DOI]
(I p)
Jennifer R Juarez, William Margolin
Irresistible curves.
EMBO J: 2009, 28(15);2147-8
[PubMed:19654604]
[WorldCat.org]
[DOI]
(I p)
Original Publications
Kenneth Briley, Peter Prepiak, Miguel J Dias, Jeanette Hahn, David Dubnau
Maf acts downstream of ComGA to arrest cell division in competent cells of B. subtilis.
Mol Microbiol: 2011, 81(1);23-39
[PubMed:21564336]
[WorldCat.org]
[DOI]
(I p)
Maria A Oliva, Sven Halbedel, Stefan M Freund, Pavel Dutow, Thomas A Leonard, Dmitry B Veprintsev, Leendert W Hamoen, Jan Löwe
Features critical for membrane binding revealed by DivIVA crystal structure.
EMBO J: 2010, 29(12);1988-2001
[PubMed:20502438]
[WorldCat.org]
[DOI]
(I p)
Suey van Baarle, Marc Bramkamp
The MinCDJ system in Bacillus subtilis prevents minicell formation by promoting divisome disassembly.
PLoS One: 2010, 5(3);e9850
[PubMed:20352045]
[WorldCat.org]
[DOI]
(I e)
Kumaran S Ramamurthi, Richard Losick
Negative membrane curvature as a cue for subcellular localization of a bacterial protein.
Proc Natl Acad Sci U S A: 2009, 106(32);13541-5
[PubMed:19666580]
[WorldCat.org]
[DOI]
(I p)
Jennifer R Juarez, William Margolin
Irresistible curves.
EMBO J: 2009, 28(15);2147-8
[PubMed:19654604]
[WorldCat.org]
[DOI]
(I p)
Rok Lenarcic, Sven Halbedel, Loek Visser, Michael Shaw, Ling Juan Wu, Jeff Errington, Davide Marenduzzo, Leendert W Hamoen
Localisation of DivIVA by targeting to negatively curved membranes.
EMBO J: 2009, 28(15);2272-82
[PubMed:19478798]
[WorldCat.org]
[DOI]
(I p)
Pamela Gamba, Jan-Willem Veening, Nigel J Saunders, Leendert W Hamoen, Richard A Daniel
Two-step assembly dynamics of the Bacillus subtilis divisome.
J Bacteriol: 2009, 191(13);4186-94
[PubMed:19429628]
[WorldCat.org]
[DOI]
(I p)
Marc Bramkamp, Robyn Emmins, Louise Weston, Catriona Donovan, Richard A Daniel, Jeff Errington
A novel component of the division-site selection system of Bacillus subtilis and a new mode of action for the division inhibitor MinCD.
Mol Microbiol: 2008, 70(6);1556-69
[PubMed:19019154]
[WorldCat.org]
[DOI]
(I p)
S E Perry, D H Edwards
Identification of a polar targeting determinant for Bacillus subtilis DivIVA.
Mol Microbiol: 2004, 54(5);1237-49
[PubMed:15554965]
[WorldCat.org]
[DOI]
(P p)
Virginie Molle, Masaya Fujita, Shane T Jensen, Patrick Eichenberger, José E González-Pastor, Jun S Liu, Richard Losick
The Spo0A regulon of Bacillus subtilis.
Mol Microbiol: 2003, 50(5);1683-701
[PubMed:14651647]
[WorldCat.org]
[DOI]
(P p)
Leendert W Hamoen, Jeffery Errington
Polar targeting of DivIVA in Bacillus subtilis is not directly dependent on FtsZ or PBP 2B.
J Bacteriol: 2003, 185(2);693-7
[PubMed:12511520]
[WorldCat.org]
[DOI]
(P p)
Frederico J Gueiros-Filho, Richard Losick
A widely conserved bacterial cell division protein that promotes assembly of the tubulin-like protein FtsZ.
Genes Dev: 2002, 16(19);2544-56
[PubMed:12368265]
[WorldCat.org]
[DOI]
(P p)
M E Karoui, J Errington
Isolation and characterization of topological specificity mutants of minD in Bacillus subtilis.
Mol Microbiol: 2001, 42(5);1211-21
[PubMed:11886553]
[WorldCat.org]
[DOI]
(P p)
H B Thomaides, M Freeman, M El Karoui, J Errington
Division site selection protein DivIVA of Bacillus subtilis has a second distinct function in chromosome segregation during sporulation.
Genes Dev: 2001, 15(13);1662-73
[PubMed:11445541]
[WorldCat.org]
[DOI]
(P p)
D H Edwards, H B Thomaides, J Errington
Promiscuous targeting of Bacillus subtilis cell division protein DivIVA to division sites in Escherichia coli and fission yeast.
EMBO J: 2000, 19(11);2719-27
[PubMed:10835369]
[WorldCat.org]
[DOI]
(P p)
D H Edwards, J Errington
The Bacillus subtilis DivIVA protein targets to the division septum and controls the site specificity of cell division.
Mol Microbiol: 1997, 24(5);905-15
[PubMed:9219999]
[WorldCat.org]
[DOI]
(P p)
J H Cha, G C Stewart
The divIVA minicell locus of Bacillus subtilis.
J Bacteriol: 1997, 179(5);1671-83
[PubMed:9045828]
[WorldCat.org]
[DOI]
(P p)