Difference between revisions of "GlcT"
(→Original publications) |
(→Original publications) |
||
| Line 136: | Line 136: | ||
<pubmed> 9663674 18086213 </pubmed> | <pubmed> 9663674 18086213 </pubmed> | ||
==Original publications== | ==Original publications== | ||
| − | <pubmed>11902727 9765562 12437213 10543968 17074746 15155854 14527945 , 19684596 | + | <pubmed>11902727 9765562 12437213 10543968 17074746 15155854 14527945 , 19684596 </pubmed> |
[[Category:Protein-coding genes]] | [[Category:Protein-coding genes]] | ||
Revision as of 10:28, 25 October 2010
- Description: Transcriptional antiterminator , controls expression of the ptsG-ptsH-ptsI operon
| Gene name | glcT |
| Synonyms | ykwA |
| Essential | no |
| Product | transcriptional antiterminator of the ptsG-ptsH-ptsI operon |
| Function | control of glucose uptake |
| Metabolic function and regulation of this protein in SubtiPathways: Central C-metabolism | |
| MW, pI | 33,0 kDa, 7.01 |
| Gene length, protein length | 855 bp, 285 amino acids |
| Immediate neighbours | ykvZ, ptsG |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 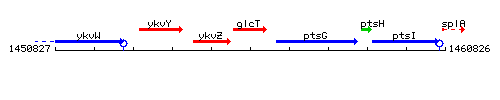
This image was kindly provided by SubtiList
| |
Contents
The gene
Basic information
- Locus tag: BSU13880
Phenotypes of a mutant
Database entries
- DBTBS entry: no entry
- SubtiList entry: [1]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: transcription antiterminator , RNA-binding protein, binds the ptsG RAT sequence
- Protein family: transcription antiterminator of the BglG/ SacY family
Extended information on the protein
- Kinetic information:
- Domains:
- RNA-binding domain (N-terminal, constitutive antiterminator)
- 2x PTS regulation domains (PRDs) (C-terminal, neg. regulated by PtsG)
- Modification: phosphorylation (His104)
- Cofactor(s):
- Effectors of protein activity:
- Localization:
Database entries
- UniProt: O31691
- KEGG entry: [2]
Additional information
Expression and regulation
- Operon:
- Sigma factor:
- Regulation:
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant: available in Stülke lab:
- Expression vector:
- pGP124 (full length, in pWH844), available in Stülke lab
- pGP114 (amino acids 1-60, RNA-binding domain, in pWH844), available in Stülke lab
- pGP230 (amino acids 1-60, RNA-binding domain with thrombin cleavage site, in pWH844), available in Stülke lab
- pGP164 (both PRDs, in pWH844), in addition diverse expression vectors for phosphorylation site mutants and for RBD mutants (all in pWH844), available in Stülke lab
- pGP424 (PRDI, in pWH844), available in Stülke lab
- pGP425 (PRDII, in pWH844), available in Stülke lab
- pGP442 (PRDI, in pGP570, with thrombin cleavage site), available in Stülke lab
- pGP443 (PRDII, in pGP570, with thrombin cleavage site), available in Stülke lab
- pGP575 (amino acids 1-60, RNA-binding domain with Strep-tag, in pGP574), available in Stülke lab
- lacZ fusion:
- GFP fusion:
- Antibody:
Labs working on this gene/protein
Jörg Stülke, University of Göttingen, Germany Homepage
Your additional remarks
References
Reviews
Fabian M Commichau, Jörg Stülke
Trigger enzymes: bifunctional proteins active in metabolism and in controlling gene expression.
Mol Microbiol: 2008, 67(4);692-702
[PubMed:18086213]
[WorldCat.org]
[DOI]
(P p)
J Stülke, M Arnaud, G Rapoport, I Martin-Verstraete
PRD--a protein domain involved in PTS-dependent induction and carbon catabolite repression of catabolic operons in bacteria.
Mol Microbiol: 1998, 28(5);865-74
[PubMed:9663674]
[WorldCat.org]
[DOI]
(P p)
Original publications