Difference between revisions of "Sandbox"
| Line 1: | Line 1: | ||
| − | * '''Description:''' | + | * '''Description:''' penicillin-binding protein 5*, D-alanyl-D-alanine carboxypeptidase <br/><br/> |
{| align="right" border="1" cellpadding="2" | {| align="right" border="1" cellpadding="2" | ||
|- | |- | ||
|style="background:#ABCDEF;" align="center"|'''Gene name''' | |style="background:#ABCDEF;" align="center"|'''Gene name''' | ||
| − | |'' | + | |''dacA'' |
|- | |- | ||
| − | |style="background:#ABCDEF;" align="center"| '''Synonyms''' || '' | + | |style="background:#ABCDEF;" align="center"| '''Synonyms''' || '' '' |
|- | |- | ||
|style="background:#ABCDEF;" align="center"| '''Essential''' || no | |style="background:#ABCDEF;" align="center"| '''Essential''' || no | ||
|- | |- | ||
| − | |style="background:#ABCDEF;" align="center"| '''Product''' || | + | |style="background:#ABCDEF;" align="center"| '''Product''' || penicillin-binding protein 5*, <br/>D-alanyl-D-alanine carboxypeptidase |
|- | |- | ||
| − | |style="background:#ABCDEF;" align="center"|'''Function''' || | + | |style="background:#ABCDEF;" align="center"|'''Function''' || carboxypeptidase |
|- | |- | ||
| − | |style="background:#ABCDEF;" align="center"| '''MW, pI''' || | + | |style="background:#ABCDEF;" align="center"| '''MW, pI''' || 48 kDa, 5.668 |
|- | |- | ||
| − | |style="background:#ABCDEF;" align="center"| '''Gene length, protein length''' || | + | |style="background:#ABCDEF;" align="center"| '''Gene length, protein length''' || 1329 bp, 443 aa |
|- | |- | ||
| − | |style="background:#ABCDEF;" align="center"|'''Immediate neighbours''' || ''[[ | + | |style="background:#ABCDEF;" align="center"|'''Immediate neighbours''' || ''[[guaB]]'', ''[[pdxS]]'' |
|- | |- | ||
|colspan="2" style="background:#FAF8CC;" align="center"|'''Hier soll was neues rein''' | |colspan="2" style="background:#FAF8CC;" align="center"|'''Hier soll was neues rein''' | ||
|- | |- | ||
| − | |colspan="2" | '''Genetic context''' <br/> [[Image: | + | |colspan="2" | '''Genetic context''' <br/> [[Image:dacA_context.gif]] |
<div align="right"> <small>This image was kindly provided by [http://genolist.pasteur.fr/SubtiList/ SubtiList]</small></div> | <div align="right"> <small>This image was kindly provided by [http://genolist.pasteur.fr/SubtiList/ SubtiList]</small></div> | ||
|- | |- | ||
| Line 29: | Line 29: | ||
__TOC__ | __TOC__ | ||
| − | <br/><br/> | + | <br/><br/><br/> |
=The gene= | =The gene= | ||
| Line 38: | Line 38: | ||
===Phenotypes of a mutant === | ===Phenotypes of a mutant === | ||
| − | |||
| − | |||
=== Database entries === | === Database entries === | ||
| − | * '''DBTBS entry:''' | + | * '''DBTBS entry:''' no entry |
| − | * '''SubtiList entry:''' [http://genolist.pasteur.fr/SubtiList/genome.cgi?gene_detail+ | + | * '''SubtiList entry:''' [http://genolist.pasteur.fr/SubtiList/genome.cgi?gene_detail+BG10074] |
=== Additional information=== | === Additional information=== | ||
| Line 66: | Line 64: | ||
* '''Domains:''' | * '''Domains:''' | ||
| − | * '''Modification:''' | + | * '''Modification:''' |
* '''Cofactor(s):''' | * '''Cofactor(s):''' | ||
| Line 72: | Line 70: | ||
* '''Effectors of protein activity:''' | * '''Effectors of protein activity:''' | ||
| − | * '''Interactions:''' | + | * '''Interactions:''' |
| − | * '''Localization:''' | + | * '''Localization:''' membrane associated [http://www.ncbi.nlm.nih.gov/pubmed/18763711 PubMed] |
=== Database entries === | === Database entries === | ||
| − | * '''Structure:''' | + | * '''Structure:''' |
* '''Swiss prot entry:''' | * '''Swiss prot entry:''' | ||
| − | * '''KEGG entry:''' [http://www.genome.jp/dbget-bin/www_bget?bsu+ | + | * '''KEGG entry:''' [http://www.genome.jp/dbget-bin/www_bget?bsu+BSU00100] |
* '''E.C. number:''' | * '''E.C. number:''' | ||
| Line 90: | Line 88: | ||
=Expression and regulation= | =Expression and regulation= | ||
| − | * '''Operon:' | + | * '''Operon:''' |
* '''[[Sigma factor]]:''' | * '''[[Sigma factor]]:''' | ||
| − | * '''Regulation:''' | + | * '''Regulation:''' |
* '''Regulatory mechanism:''' | * '''Regulatory mechanism:''' | ||
| − | * '''Additional information:''' | + | * '''Additional information:''' |
=Biological materials = | =Biological materials = | ||
| Line 120: | Line 118: | ||
=References= | =References= | ||
| − | # | + | # Hahne et al. (2008) From complementarity to comprehensiveness - targeting the membrane proteome of growing ''Bacillus subtilis'' by divergent approaches. Proteomics '''8:''' 4123-4136 [http://www.ncbi.nlm.nih.gov/pubmed/18763711 PubMed] |
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
Revision as of 20:44, 15 April 2009
- Description: penicillin-binding protein 5*, D-alanyl-D-alanine carboxypeptidase
| Gene name | dacA |
| Synonyms | |
| Essential | no |
| Product | penicillin-binding protein 5*, D-alanyl-D-alanine carboxypeptidase |
| Function | carboxypeptidase |
| MW, pI | 48 kDa, 5.668 |
| Gene length, protein length | 1329 bp, 443 aa |
| Immediate neighbours | guaB, pdxS |
| Hier soll was neues rein | |
Genetic context 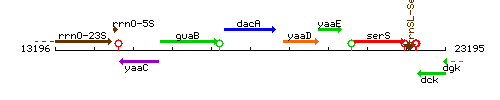
This image was kindly provided by SubtiList
| |
Contents
The gene
Basic information
- Coordinates:
Phenotypes of a mutant
Database entries
- DBTBS entry: no entry
- SubtiList entry: [1]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family:
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Interactions:
- Localization: membrane associated PubMed
Database entries
- Structure:
- Swiss prot entry:
- KEGG entry: [2]
- E.C. number:
Additional information
Expression and regulation
- Operon:
- Regulation:
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
- Hahne et al. (2008) From complementarity to comprehensiveness - targeting the membrane proteome of growing Bacillus subtilis by divergent approaches. Proteomics 8: 4123-4136 PubMed