Difference between revisions of "MurE"
| Line 128: | Line 128: | ||
* '''Additional information:''' | * '''Additional information:''' | ||
| + | ** number of protein molecules per cell (minimal medium with glucose and ammonium): 297 {{PubMed|24696501}} | ||
| + | ** number of protein molecules per cell (complex medium with amino acids, without glucose): 1548 {{PubMed|24696501}} | ||
=Biological materials = | =Biological materials = | ||
Revision as of 10:51, 17 April 2014
- Description: UDP-N-acetylmuramoyl-L-alanyl-D-glutamyl-meso-2,6-diaminopimelate synthetase
| Gene name | murE |
| Synonyms | |
| Essential | yes PubMed |
| Product | UDP-N-acetylmuramoyl-L-alanyl-D-glutamyl- meso-2,6-diaminopimelate synthetase |
| Function | peptidoglycan precursor biosynthesis |
| Gene expression levels in SubtiExpress: murE | |
| Metabolic function and regulation of this protein in SubtiPathways: murE | |
| MW, pI | 54 kDa, 5.558 |
| Gene length, protein length | 1482 bp, 494 aa |
| Immediate neighbours | spoVD, mraY |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 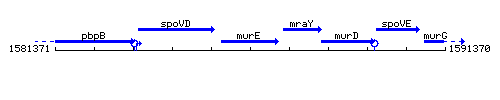
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed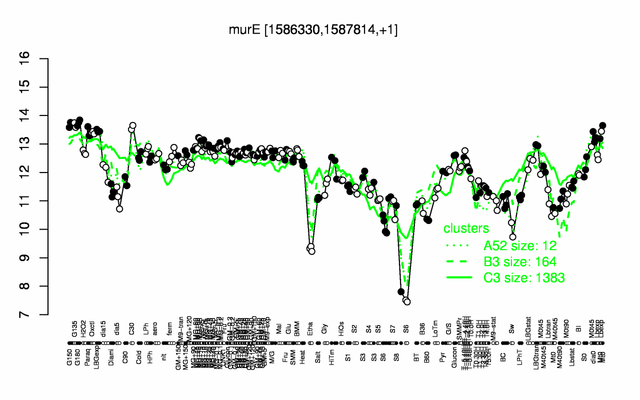
| |
Contents
Categories containing this gene/protein
cell wall synthesis, biosynthesis of cell wall components, essential genes
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU15180
Phenotypes of a mutant
essential PubMed
Database entries
- BsubCyc: BSU15180
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: ATP + UDP-N-acetylmuramoyl-L-alanyl-D-glutamate + meso-2,6-diaminoheptanedioate = ADP + phosphate + UDP-N-acetylmuramoyl-L-alanyl-D-gamma-glutamyl-meso-2,6-diamino-heptanedioate (according to Swiss-Prot)
- Protein family: MurE subfamily (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Localization:
- cytoplasm (according to Swiss-Prot)
Database entries
- BsubCyc: BSU15180
- UniProt: Q03523
- KEGG entry: [3]
- E.C. number: 6.3.2.13
Additional information
Expression and regulation
- Sigma factor:
- Regulation:
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References