Difference between revisions of "YtsJ"
| Line 143: | Line 143: | ||
=References= | =References= | ||
| − | + | <pubmed>22740702</pubmed> | |
| − | <pubmed>16479537, 16788182 ,12949160 9387221 | + | <pubmed>16479537, 16788182 ,12949160 9387221</pubmed> |
[[Category:Protein-coding genes]] | [[Category:Protein-coding genes]] | ||
Revision as of 17:27, 30 October 2012
- Description: malic enzyme, forms a transhydrogenation cycle with MalS for balancing of NADPH
| Gene name | ytsJ |
| Synonyms | |
| Essential | no |
| Product | NADP-dependent malate dehydrogenase |
| Function | malate utilization |
| Gene expression levels in SubtiExpress: ytsJ | |
| Metabolic function and regulation of this protein in SubtiPathways: Central C-metabolism | |
| MW, pI | 43 kDa, 5.046 |
| Gene length, protein length | 1230 bp, 410 aa |
| Immediate neighbours | accD, dnaE |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 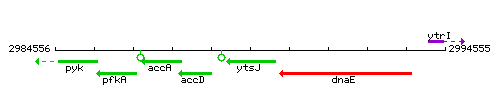
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed
| |
Contents
Categories containing this gene/protein
utilization of specific carbon sources
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU29220
Phenotypes of a mutant
Poor growth with malate as single carbon source PubMed
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: (S)-malate + NADP+ = pyruvate + CO2 + NADPH (according to Swiss-Prot) malate <--> pyruvate
- Protein family: malic enzymes family (according to Swiss-Prot)
- Paralogous protein(s): MleA
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Interactions:
Database entries
- Structure: 1WW8 (from Pyrococcus horikoshii, 46% identity, 63% similarity)
- UniProt: O34962
- KEGG entry: [3]
- E.C. number: 1.1.1.38
Additional information
Expression and regulation
- Sigma factor:
- Regulation:
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant:
- GP612 (spc), available in Jörg Stülke's lab
- GP1143 (spc), available in Jörg Stülke's lab
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- FLAG-tag construct: GP1131 (spc, based on pGP1331), available in Jörg Stülke's lab
- Antibody:
Labs working on this gene/protein
Stephane Aymerich, Microbiology and Molecular Genetics, INRA Paris-Grignon, France
Your additional remarks
References
Guillaume Lerondel, Thierry Doan, Nicola Zamboni, Uwe Sauer, Stéphane Aymerich
YtsJ has the major physiological role of the four paralogous malic enzyme isoforms in Bacillus subtilis.
J Bacteriol: 2006, 188(13);4727-36
[PubMed:16788182]
[WorldCat.org]
[DOI]
(P p)
Jean-Christophe Meile, Ling Juan Wu, S Dusko Ehrlich, Jeff Errington, Philippe Noirot
Systematic localisation of proteins fused to the green fluorescent protein in Bacillus subtilis: identification of new proteins at the DNA replication factory.
Proteomics: 2006, 6(7);2135-46
[PubMed:16479537]
[WorldCat.org]
[DOI]
(P p)
Thierry Doan, Pascale Servant, Shigeo Tojo, Hirotake Yamaguchi, Guillaume Lerondel, Ken-Ichi Yoshida, Yasutaro Fujita, Stéphane Aymerich
The Bacillus subtilis ywkA gene encodes a malic enzyme and its transcription is activated by the YufL/YufM two-component system in response to malate.
Microbiology (Reading): 2003, 149(Pt 9);2331-2343
[PubMed:12949160]
[WorldCat.org]
[DOI]
(P p)
Alia Lapidus, Nathalie Galleron, Alexei Sorokin, S Dusko Ehrlich
Sequencing and functional annotation of the Bacillus subtilis genes in the 200 kb rrnB-dnaB region.
Microbiology (Reading): 1997, 143 ( Pt 11);3431-3441
[PubMed:9387221]
[WorldCat.org]
[DOI]
(P p)