Difference between revisions of "MurB"
| Line 14: | Line 14: | ||
|style="background:#ABCDEF;" align="center"|'''Function''' || peptidoglycan precursor biosynthesis | |style="background:#ABCDEF;" align="center"|'''Function''' || peptidoglycan precursor biosynthesis | ||
|- | |- | ||
| − | |colspan="2" style="background:#FAF8CC;" align="center"| '''Gene expression levels in [http:// | + | |colspan="2" style="background:#FAF8CC;" align="center"| '''Gene expression levels in [http://subtiwiki.uni-goettingen.de/apps/expression/ ''Subti''Express]''': [http://subtiwiki.uni-goettingen.de/apps/expression/expression.php?search=BSU15230 murB] |
|- | |- | ||
|colspan="2" style="background:#FAF8CC;" align="center"| '''Metabolic function and regulation of this protein in [[SubtiPathways|''Subti''Pathways]]: <br/>[http://subtiwiki.uni-goettingen.de/pathways/cellwall.html Cell wall]''' | |colspan="2" style="background:#FAF8CC;" align="center"| '''Metabolic function and regulation of this protein in [[SubtiPathways|''Subti''Pathways]]: <br/>[http://subtiwiki.uni-goettingen.de/pathways/cellwall.html Cell wall]''' | ||
| Line 24: | Line 24: | ||
|style="background:#ABCDEF;" align="center"|'''Immediate neighbours''' || ''[[murG]]'', ''[[divIB]]'' | |style="background:#ABCDEF;" align="center"|'''Immediate neighbours''' || ''[[murG]]'', ''[[divIB]]'' | ||
|- | |- | ||
| − | | | + | |style="background:#FAF8CC;" align="center"|'''Sequences'''||[http://bsubcyc.org/BSUB/sequence-aa?type=GENE&object=BSU15230 Protein] [http://bsubcyc.org/BSUB/sequence?type=GENE&object=BSU15230 DNA] [http://bsubcyc.org/BSUB/seq-selector?chromosome=CHROM-1&object=BSU15230 Advanced_DNA] |
|- | |- | ||
|colspan="2" | '''Genetic context''' <br/> [[Image:murB_context.gif]] | |colspan="2" | '''Genetic context''' <br/> [[Image:murB_context.gif]] | ||
Revision as of 13:45, 13 May 2013
- Description: UDP-N-acetylenolpyruvoylglucosamine reductase
| Gene name | murB |
| Synonyms | ylxC |
| Essential | yes PubMed |
| Product | UDP-N-acetylenolpyruvoylglucosamine reductase |
| Function | peptidoglycan precursor biosynthesis |
| Gene expression levels in SubtiExpress: murB | |
| Metabolic function and regulation of this protein in SubtiPathways: Cell wall | |
| MW, pI | 32 kDa, 6.087 |
| Gene length, protein length | 909 bp, 303 aa |
| Immediate neighbours | murG, divIB |
| Sequences | Protein DNA Advanced_DNA |
Genetic context 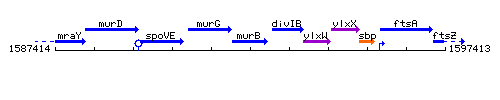
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed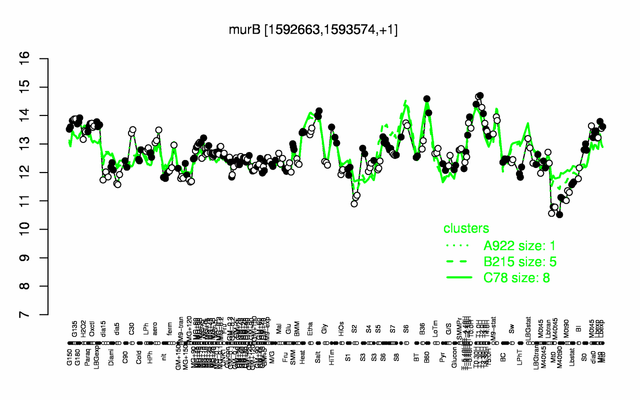
| |
Contents
Categories containing this gene/protein
cell wall synthesis, biosynthesis of cell wall components, sporulation proteins, cell envelope stress proteins (controlled by SigM, V, W, X, Y), essential genes
This gene is a member of the following regulons
SigE regulon, SigM regulon, SpoIIID regulon
The gene
Basic information
- Locus tag: BSU15230
Phenotypes of a mutant
essential PubMed
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: UDP-N-acetylmuramate + NADP+ = UDP-N-acetyl-3-O-(1-carboxyvinyl)-D-glucosamine + NADPH (according to Swiss-Prot)
- Protein family: FAD-binding PCMH-type domain (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Localization: cytoplasm (according to Swiss-Prot)
Database entries
- UniProt: P18579
- KEGG entry: [3]
- E.C. number: 1.1.1.158
Additional information
Expression and regulation
- Operon:
- Regulation:
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Adriano Henriques, Lisbon, Portugal homepage
Your additional remarks
References