Difference between revisions of "CwlS"
(→Extended information on the protein) |
|||
| Line 67: | Line 67: | ||
* '''Protein family:''' Cu-Zn superoxide dismutase family (according to Swiss-Prot) | * '''Protein family:''' Cu-Zn superoxide dismutase family (according to Swiss-Prot) | ||
| − | * '''Paralogous protein(s):''' | + | * '''Paralogous protein(s):''' the C-terminal D,L-endopeptidase domains of [[LytE]], [[LytF]], [[CwlS]], and [[CwlO]] exhibit strong sequence similarity |
=== Extended information on the protein === | === Extended information on the protein === | ||
| Line 74: | Line 74: | ||
* '''Domains:''' | * '''Domains:''' | ||
| + | ** C-terminal D,L-endopeptidase domain {{PubMed|22139507}} | ||
* '''Modification:''' | * '''Modification:''' | ||
| Line 85: | Line 86: | ||
* '''[[Localization]]:''' | * '''[[Localization]]:''' | ||
** cell membrane (according to Swiss-Prot) | ** cell membrane (according to Swiss-Prot) | ||
| + | ** localizes to cell septa and poles {{PubMed|22139507}} | ||
=== Database entries === | === Database entries === | ||
| Line 102: | Line 104: | ||
* '''Operon:''' | * '''Operon:''' | ||
| − | * '''[[Sigma factor]]:''' | + | * '''[[Sigma factor]]:''' [[SigD]], [[SigH]], according to {{PubMed|22139507}} |
* '''Regulation:''' | * '''Regulation:''' | ||
| Line 133: | Line 135: | ||
=References= | =References= | ||
'''Additional publications:''' {{PubMed|20817675}} | '''Additional publications:''' {{PubMed|20817675}} | ||
| − | <pubmed>16855244,12850135, </pubmed> | + | <pubmed>16855244,12850135, 22139507 </pubmed> |
[[Category:Protein-coding genes]] | [[Category:Protein-coding genes]] | ||
Revision as of 13:00, 8 December 2011
- Description: D,L-endopeptidase, peptidoglycan hydrolase
| Gene name | cwlS |
| Synonyms | yojL |
| Essential | no |
| Product | D,L-endopeptidase, peptidoglycan hydrolase |
| Function | cell wall metabolism |
| MW, pI | 44 kDa, 10.438 |
| Gene length, protein length | 1242 bp, 414 aa |
| Immediate neighbours | yojM, yojK |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 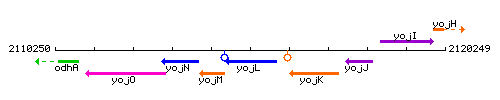
This image was kindly provided by SubtiList
| |
Contents
Categories containing this gene/protein
cell wall degradation/ turnover, membrane proteins
This gene is a member of the following regulons
Abh regulon, AbrB regulon, CcpA regulon
The gene
Basic information
- Locus tag: BSU19410
Phenotypes of a mutant
Database entries
- DBTBS entry: no entry
- SubtiList entry: [1]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: Hydrolysis of gamma-D-glutamyl bonds to the L-terminus (position 7) of meso-diaminopimelic acid (meso-A2pm) in 7-(L-Ala-gamma-D-Glu)-meso-A2pm and 7-(L-Ala-gamma-D-Glu)-7-(D-Ala)-meso-A2pm (according to Swiss-Prot)
- Protein family: Cu-Zn superoxide dismutase family (according to Swiss-Prot)
- Paralogous protein(s): the C-terminal D,L-endopeptidase domains of LytE, LytF, CwlS, and CwlO exhibit strong sequence similarity
Extended information on the protein
- Kinetic information:
- Domains:
- C-terminal D,L-endopeptidase domain PubMed
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Localization:
- cell membrane (according to Swiss-Prot)
- localizes to cell septa and poles PubMed
Database entries
- Structure:
- UniProt: O31852
- KEGG entry: [2]
- E.C. number:
Additional information
Expression and regulation
- Operon:
- Sigma factor: SigD, SigH, according to PubMed
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Additional publications: PubMed