Difference between revisions of "TsaC"
| Line 1: | Line 1: | ||
| − | * '''Description:''' | + | * '''Description:''' biosynthesis of the hypermodified base threonylcarbamoyladenosine (t(6)A) <br/><br/> |
{| align="right" border="1" cellpadding="2" | {| align="right" border="1" cellpadding="2" | ||
| Line 12: | Line 12: | ||
|style="background:#ABCDEF;" align="center"| '''Product''' || unknown | |style="background:#ABCDEF;" align="center"| '''Product''' || unknown | ||
|- | |- | ||
| − | |style="background:#ABCDEF;" align="center"|'''Function''' || | + | |style="background:#ABCDEF;" align="center"|'''Function''' || tRNA modification |
|- | |- | ||
|style="background:#ABCDEF;" align="center"| '''MW, pI''' || 36 kDa, 5.82 | |style="background:#ABCDEF;" align="center"| '''MW, pI''' || 36 kDa, 5.82 | ||
| Line 52: | Line 52: | ||
=== Basic information/ Evolution === | === Basic information/ Evolution === | ||
| − | * '''Catalyzed reaction/ biological activity:''' | + | * '''Catalyzed reaction/ biological activity:''' biosynthesis of the hypermodified base threonylcarbamoyladenosine (t(6)A), a universal modification found at position 37 of ANN decoding tRNAs, which imparts a unique structure to the anticodon loop enhancing its binding to ribosomes in vitro (shown for the corresponding ''E. coli'' protein YrdC) {{PubMed|19287007}} |
* '''Protein family:''' SUA5 family (according to Swiss-Prot) | * '''Protein family:''' SUA5 family (according to Swiss-Prot) | ||
| Line 76: | Line 76: | ||
=== Database entries === | === Database entries === | ||
| − | * '''Structure:''' | + | * '''Structure:''' [http://www.rcsb.org/pdb/explore/explore.do?pdbId=1HRU 1HRU] (the homolog from ''E. coli'') {{PubMed|19287007}} |
* '''UniProt:''' [http://www.uniprot.org/uniprot/P39153 P39153] | * '''UniProt:''' [http://www.uniprot.org/uniprot/P39153 P39153] | ||
| Line 119: | Line 119: | ||
=References= | =References= | ||
| − | <pubmed>17005971 17932079</pubmed> | + | <pubmed>17005971 17932079 19287007 </pubmed> |
[[Category:Protein-coding genes]] | [[Category:Protein-coding genes]] | ||
Revision as of 12:30, 15 December 2009
- Description: biosynthesis of the hypermodified base threonylcarbamoyladenosine (t(6)A)
| Gene name | ywlC |
| Synonyms | ipc-29d |
| Essential | no |
| Product | unknown |
| Function | tRNA modification |
| MW, pI | 36 kDa, 5.82 |
| Gene length, protein length | 1038 bp, 346 aa |
| Immediate neighbours | ywlD, ywlB |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 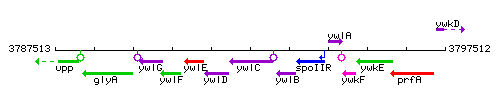
This image was kindly provided by SubtiList
| |
Contents
The gene
Basic information
- Locus tag: BSU36950
Phenotypes of a mutant
Database entries
- DBTBS entry: no entry
- SubtiList entry: [1]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: biosynthesis of the hypermodified base threonylcarbamoyladenosine (t(6)A), a universal modification found at position 37 of ANN decoding tRNAs, which imparts a unique structure to the anticodon loop enhancing its binding to ribosomes in vitro (shown for the corresponding E. coli protein YrdC) PubMed
- Protein family: SUA5 family (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Interactions:
- Localization:
Database entries
- UniProt: P39153
- KEGG entry: [2]
- E.C. number:
Additional information
Expression and regulation
- Operon:
- Regulation:
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Basma El Yacoubi, Benjamin Lyons, Yulien Cruz, Robert Reddy, Brian Nordin, Fabio Agnelli, James R Williamson, Paul Schimmel, Manal A Swairjo, Valérie de Crécy-Lagard
The universal YrdC/Sua5 family is required for the formation of threonylcarbamoyladenosine in tRNA.
Nucleic Acids Res: 2009, 37(9);2894-909
[PubMed:19287007]
[WorldCat.org]
[DOI]
(I p)
Shu Ishikawa, Yoshitoshi Ogura, Mika Yoshimura, Hajime Okumura, Eunha Cho, Yoshikazu Kawai, Ken Kurokawa, Taku Oshima, Naotake Ogasawara
Distribution of stable DnaA-binding sites on the Bacillus subtilis genome detected using a modified ChIP-chip method.
DNA Res: 2007, 14(4);155-68
[PubMed:17932079]
[WorldCat.org]
[DOI]
(P p)
Alison Hunt, Joy P Rawlins, Helena B Thomaides, Jeff Errington
Functional analysis of 11 putative essential genes in Bacillus subtilis.
Microbiology (Reading): 2006, 152(Pt 10);2895-2907
[PubMed:17005971]
[WorldCat.org]
[DOI]
(P p)