Difference between revisions of "IseA"
| (37 intermediate revisions by 5 users not shown) | |||
| Line 1: | Line 1: | ||
| − | * '''Description:''' inhibits in vitro activity of cell wall endopeptidases LytE and LytF, inhibits cell separation <br/><br/> | + | * '''Description:''' inhibits in vitro activity of cell wall endopeptidases [[LytE]] and [[LytF]], inhibits cell separation <br/><br/> |
{| align="right" border="1" cellpadding="2" | {| align="right" border="1" cellpadding="2" | ||
| Line 13: | Line 13: | ||
|- | |- | ||
|style="background:#ABCDEF;" align="center"|'''Function''' || protection against cell envelope stress | |style="background:#ABCDEF;" align="center"|'''Function''' || protection against cell envelope stress | ||
| + | |- | ||
| + | |colspan="2" style="background:#FAF8CC;" align="center"| '''Gene expression levels in [http://subtiwiki.uni-goettingen.de/apps/expression/ ''Subti''Express]''': [http://subtiwiki.uni-goettingen.de/apps/expression/expression.php?search=BSU18380 iseA] | ||
| + | |- | ||
| + | |colspan="2" style="background:#FAF8CC;" align="center"| '''Interactions involving this protein in [http://subtiwiki.uni-goettingen.de/interact/ ''Subt''Interact]''': [http://subtiwiki.uni-goettingen.de/interact/index.php?protein=IseA IseA] | ||
|- | |- | ||
|style="background:#ABCDEF;" align="center"| '''MW, pI''' || 19 kDa, 10.176 | |style="background:#ABCDEF;" align="center"| '''MW, pI''' || 19 kDa, 10.176 | ||
| Line 20: | Line 24: | ||
|style="background:#ABCDEF;" align="center"|'''Immediate neighbours''' || ''[[yoeA]]'', ''[[trnSL-Arg1]]'' | |style="background:#ABCDEF;" align="center"|'''Immediate neighbours''' || ''[[yoeA]]'', ''[[trnSL-Arg1]]'' | ||
|- | |- | ||
| − | | | + | |style="background:#FAF8CC;" align="center"|'''Sequences'''||[http://bsubcyc.org/BSUB/sequence-aa?type=GENE&object=BSU18380 Protein] [http://bsubcyc.org/BSUB/sequence?type=GENE&object=BSU18380 DNA] [http://bsubcyc.org/BSUB/seq-selector?chromosome=CHROM-1&object=BSU18380 DNA_with_flanks] |
|- | |- | ||
|- | |- | ||
| Line 26: | Line 30: | ||
|colspan="2" | '''Genetic context''' <br/> [[Image:yoeB_context.gif]] | |colspan="2" | '''Genetic context''' <br/> [[Image:yoeB_context.gif]] | ||
<div align="right"> <small>This image was kindly provided by [http://genolist.pasteur.fr/SubtiList/ SubtiList]</small></div> | <div align="right"> <small>This image was kindly provided by [http://genolist.pasteur.fr/SubtiList/ SubtiList]</small></div> | ||
| + | |- | ||
| + | |colspan="2" |'''[http://genome.jouy.inra.fr/cgi-bin/seb/viewdetail.py?id=iseA_2002637_2003182_1 Expression at a glance]'''   {{PubMed|22383849}}<br/>[[Image:iseA_expression.png|500px|link=http://subtiwiki.uni-goettingen.de/apps/expression/expression.php?search=BSU18380]] | ||
|- | |- | ||
|} | |} | ||
__TOC__ | __TOC__ | ||
| + | <br/><br/><br/><br/> | ||
| + | <br/><br/><br/><br/> | ||
| + | <br/><br/><br/><br/> | ||
| + | <br/><br/> | ||
| − | + | = [[Categories]] containing this gene/protein = | |
| + | {{SubtiWiki category|[[cell wall degradation/ turnover]]}}, | ||
| + | {{SubtiWiki category|[[membrane proteins]]}} | ||
| + | |||
| + | = This gene is a member of the following [[regulons]] = | ||
| + | {{SubtiWiki regulon|[[WalR regulon]]}} | ||
=The gene= | =The gene= | ||
| Line 37: | Line 52: | ||
=== Basic information === | === Basic information === | ||
| − | * ''' | + | * '''Locus tag:''' BSU18380 |
===Phenotypes of a mutant === | ===Phenotypes of a mutant === | ||
=== Database entries === | === Database entries === | ||
| + | * '''BsubCyc:''' [http://bsubcyc.org/BSUB/NEW-IMAGE?type=NIL&object=BSU18380&redirect=T BSU18380] | ||
* '''DBTBS entry:''' no entry | * '''DBTBS entry:''' no entry | ||
| Line 48: | Line 64: | ||
=== Additional information=== | === Additional information=== | ||
| + | |||
| + | |||
| Line 72: | Line 90: | ||
* '''Effectors of protein activity:''' | * '''Effectors of protein activity:''' | ||
| − | * '''Interactions:''' | + | * '''[[SubtInteract|Interactions]]:''' |
| + | ** [[IseA]]-[[LytF]] (catalytic domain) {{PubMed|18761694}} | ||
| − | * '''Localization:''' | + | * '''[[Localization]]:''' cell membrane (according to Swiss-Prot), localized to cell separation sites |
=== Database entries === | === Database entries === | ||
| + | * '''BsubCyc:''' [http://bsubcyc.org/BSUB/NEW-IMAGE?type=NIL&object=BSU18380&redirect=T BSU18380] | ||
* '''Structure:''' | * '''Structure:''' | ||
| − | * ''' | + | * '''UniProt:''' [http://www.uniprot.org/uniprot/O34841 O34841] |
| − | * '''KEGG entry:''' [http://www.genome.jp/dbget-bin/www_bget?bsu | + | * '''KEGG entry:''' [http://www.genome.jp/dbget-bin/www_bget?bsu:BSU18380] |
* '''E.C. number:''' | * '''E.C. number:''' | ||
| Line 90: | Line 110: | ||
=Expression and regulation= | =Expression and regulation= | ||
| − | * '''Operon:''' | + | * '''Operon:''' ''iseA'' {{PubMed|22383849}} |
| + | |||
| + | * '''Expression browser:''' [http://genome.jouy.inra.fr/cgi-bin/seb/viewdetail.py?id=iseA_2002637_2003182_1 iseA] {{PubMed|22383849}} | ||
| − | * ''' | + | * '''Sigma factor:''' |
* '''Regulation:''' | * '''Regulation:''' | ||
| + | ** repressed by [[WalR]] {{PubMed|17581128}} | ||
| + | ** induced by vancomycin {{PubMed|17483219}} | ||
| + | ** strongly induced in response to glucose starvation in M9 medium {{PubMed|23033921}} | ||
* '''Regulatory mechanism:''' | * '''Regulatory mechanism:''' | ||
| + | ** [[WalR]]: transcription repression {{PubMed|17581128}} | ||
| − | * '''Additional information:''' | + | * '''Additional information:''' |
| + | ** number of protein molecules per cell (minimal medium with glucose and ammonium): 188 {{PubMed|24696501}} | ||
=Biological materials = | =Biological materials = | ||
| Line 119: | Line 146: | ||
=References= | =References= | ||
| + | <pubmed>17581128,18761694,17483219, 20059685,22383849 23091053,23033921 23199363</pubmed> | ||
| − | + | [[Category:Protein-coding genes]] | |
Latest revision as of 10:59, 17 April 2014
- Description: inhibits in vitro activity of cell wall endopeptidases LytE and LytF, inhibits cell separation
| Gene name | iseA |
| Synonyms | yoeB |
| Essential | no |
| Product | inhibitor of autolysins |
| Function | protection against cell envelope stress |
| Gene expression levels in SubtiExpress: iseA | |
| Interactions involving this protein in SubtInteract: IseA | |
| MW, pI | 19 kDa, 10.176 |
| Gene length, protein length | 543 bp, 181 aa |
| Immediate neighbours | yoeA, trnSL-Arg1 |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed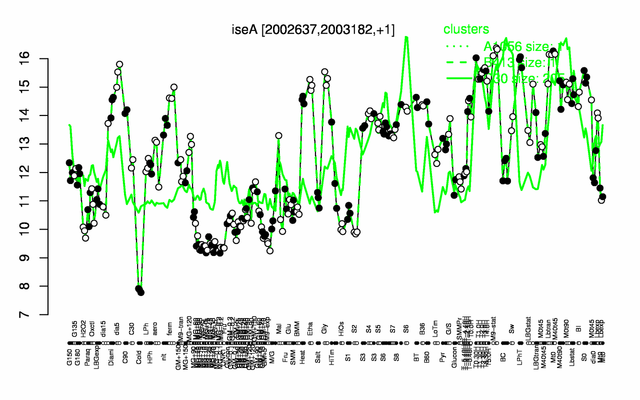
| |
Contents
Categories containing this gene/protein
cell wall degradation/ turnover, membrane proteins
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU18380
Phenotypes of a mutant
Database entries
- BsubCyc: BSU18380
- DBTBS entry: no entry
- SubtiList entry: [1]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family:
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Localization: cell membrane (according to Swiss-Prot), localized to cell separation sites
Database entries
- BsubCyc: BSU18380
- Structure:
- UniProt: O34841
- KEGG entry: [2]
- E.C. number:
Additional information
Expression and regulation
- Operon: iseA PubMed
- Sigma factor:
- Regulation:
- Additional information:
- number of protein molecules per cell (minimal medium with glucose and ammonium): 188 PubMed
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Letal I Salzberg, Leagh Powell, Karsten Hokamp, Eric Botella, David Noone, Kevin M Devine
The WalRK (YycFG) and σ(I) RsgI regulators cooperate to control CwlO and LytE expression in exponentially growing and stressed Bacillus subtilis cells.
Mol Microbiol: 2013, 87(1);180-95
[PubMed:23199363]
[WorldCat.org]
[DOI]
(I p)
Ryoichi Arai, Sadaharu Fukui, Naoya Kobayashi, Junichi Sekiguchi
Solution structure of IseA, an inhibitor protein of DL-endopeptidases from Bacillus subtilis, reveals a novel fold with a characteristic inhibitory loop.
J Biol Chem: 2012, 287(53);44736-48
[PubMed:23091053]
[WorldCat.org]
[DOI]
(I p)
Imke G de Jong, Jan-Willem Veening, Oscar P Kuipers
Single cell analysis of gene expression patterns during carbon starvation in Bacillus subtilis reveals large phenotypic variation.
Environ Microbiol: 2012, 14(12);3110-21
[PubMed:23033921]
[WorldCat.org]
[DOI]
(I p)
Pierre Nicolas, Ulrike Mäder, Etienne Dervyn, Tatiana Rochat, Aurélie Leduc, Nathalie Pigeonneau, Elena Bidnenko, Elodie Marchadier, Mark Hoebeke, Stéphane Aymerich, Dörte Becher, Paola Bisicchia, Eric Botella, Olivier Delumeau, Geoff Doherty, Emma L Denham, Mark J Fogg, Vincent Fromion, Anne Goelzer, Annette Hansen, Elisabeth Härtig, Colin R Harwood, Georg Homuth, Hanne Jarmer, Matthieu Jules, Edda Klipp, Ludovic Le Chat, François Lecointe, Peter Lewis, Wolfram Liebermeister, Anika March, Ruben A T Mars, Priyanka Nannapaneni, David Noone, Susanne Pohl, Bernd Rinn, Frank Rügheimer, Praveen K Sappa, Franck Samson, Marc Schaffer, Benno Schwikowski, Leif Steil, Jörg Stülke, Thomas Wiegert, Kevin M Devine, Anthony J Wilkinson, Jan Maarten van Dijl, Michael Hecker, Uwe Völker, Philippe Bessières, Philippe Noirot
Condition-dependent transcriptome reveals high-level regulatory architecture in Bacillus subtilis.
Science: 2012, 335(6072);1103-6
[PubMed:22383849]
[WorldCat.org]
[DOI]
(I p)
Hiroki Yamamoto, Masayuki Hashimoto, Yuhei Higashitsuji, Hiroyuki Harada, Nozomi Hariyama, Lisa Takahashi, Tomoaki Iwashita, Seika Ooiwa, Junichi Sekiguchi
Post-translational control of vegetative cell separation enzymes through a direct interaction with specific inhibitor IseA in Bacillus subtilis.
Mol Microbiol: 2008, 70(1);168-82
[PubMed:18761694]
[WorldCat.org]
[DOI]
(I p)
Paola Bisicchia, David Noone, Efthimia Lioliou, Alistair Howell, Sarah Quigley, Thomas Jensen, Hanne Jarmer, Kevin M Devine
The essential YycFG two-component system controls cell wall metabolism in Bacillus subtilis.
Mol Microbiol: 2007, 65(1);180-200
[PubMed:17581128]
[WorldCat.org]
[DOI]
(P p)
Letal I Salzberg, John D Helmann
An antibiotic-inducible cell wall-associated protein that protects Bacillus subtilis from autolysis.
J Bacteriol: 2007, 189(13);4671-80
[PubMed:17483219]
[WorldCat.org]
[DOI]
(P p)